
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,289,172 – 22,289,326 |
| Length | 154 |
| Max. P | 0.753073 |

| Location | 22,289,172 – 22,289,286 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.01 |
| Mean single sequence MFE | -31.83 |
| Consensus MFE | -25.48 |
| Energy contribution | -25.60 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.749360 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22289172 114 - 23771897 AAAUUGUUUAAACAAUAGAACAAGGUGGGCUGAGAGGCGGUCAAAAGCCGUUUCCUUUUACGAAUUCUUCGUAGAACACGGGUCGAGAAGGAGAACAGAGUUCCACA------UAAUUAA ...((((((........)))))).((((((((...(((........))).(((((((((.(((....(((((.....))))))))))))))))).)))...))))).------....... ( -29.50) >DroSec_CAF1 47590 114 - 1 AAAUUGUUUAGACAAUAGAAGAAGGUGGGCUGAGAGGCGGUCGCAAGCCGUUUCUUUUUACGAAUUCUUUGUAGAACAUGGGUCGAAAAGGAGAACAGAGUUCCAUA------UAAUUAA ..(((((....))))).((((((..((....(((((((((.(....))))))))))....))..))))))...((((.((..((........)).))..))))....------....... ( -28.60) >DroEre_CAF1 46056 120 - 1 AAAUUGUUUAAACAAUAGAAGAAGGCGGCCUGAGAGGCGGUCACAAGCCGCUUCUUUUUACGUAUUCUUCGUAGAACGUGGCUCGAAAAGGAGAACAGAGCUCUAUACGAGUGUAAUUAA ..(((((....))))).((((((.(((....((((((((((.....))))))))))....))).))))))..........(((((....((((.......))))...)))))........ ( -37.40) >consensus AAAUUGUUUAAACAAUAGAAGAAGGUGGGCUGAGAGGCGGUCACAAGCCGUUUCUUUUUACGAAUUCUUCGUAGAACAUGGGUCGAAAAGGAGAACAGAGUUCCAUA______UAAUUAA ..(((((....))))).((((((..((....((((((((((.....))))))))))....))..))))))((.((((...(.((........)).)...)))).)).............. (-25.48 = -25.60 + 0.12)

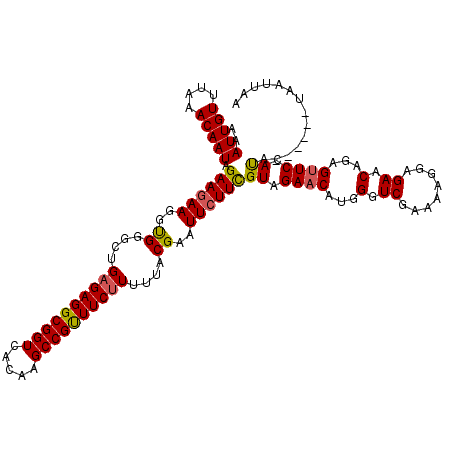

| Location | 22,289,206 – 22,289,326 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.22 |
| Mean single sequence MFE | -31.17 |
| Consensus MFE | -22.56 |
| Energy contribution | -22.57 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.753073 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 22289206 120 - 23771897 UCAUUAAAAGACAUAGGCCAGACGAAUAAGCAUUCGACAAAAAUUGUUUAAACAAUAGAACAAGGUGGGCUGAGAGGCGGUCAAAAGCCGUUUCCUUUUACGAAUUCUUCGUAGAACACG ...............((((...(((((....))))).((....((((((........))))))..))))))..((((((((.....)))))))).((((((((.....)))))))).... ( -29.20) >DroSec_CAF1 47624 106 - 1 UCAUUAGAAGACAG--------------AGCGUCCGACAAAAAUUGUUUAGACAAUAGAAGAAGGUGGGCUGAGAGGCGGUCGCAAGCCGUUUCUUUUUACGAAUUCUUUGUAGAACAUG ..............--------------.(((((.((((.....))))..)))....((((((..((....(((((((((.(....))))))))))....))..))))))))........ ( -26.00) >DroEre_CAF1 46096 119 - 1 UCAUUAGAAGACUGAGCCCAAAUG-UUGGGCACCCGACAAAAAUUGUUUAAACAAUAGAAGAAGGCGGCCUGAGAGGCGGUCACAAGCCGCUUCUUUUUACGUAUUCUUCGUAGAACGUG ...((((....))))(((((....-.)))))...((......(((((....))))).((((((.(((....((((((((((.....))))))))))....))).))))))......)).. ( -38.30) >consensus UCAUUAGAAGACAGAG_CCA_A_G__U_AGCAUCCGACAAAAAUUGUUUAAACAAUAGAAGAAGGUGGGCUGAGAGGCGGUCACAAGCCGUUUCUUUUUACGAAUUCUUCGUAGAACAUG .............................((...........(((((....))))).((((((..((....((((((((((.....))))))))))....))..))))))))........ (-22.56 = -22.57 + 0.01)

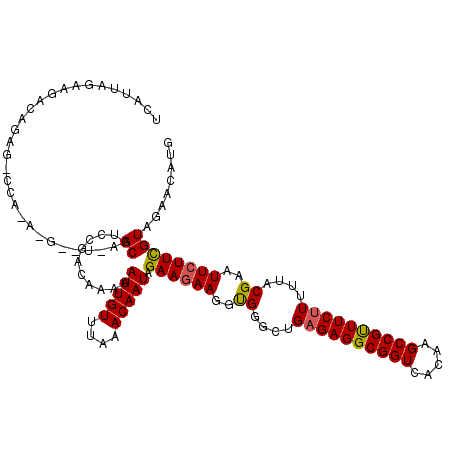

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:17:03 2006