| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,201,887 – 22,202,005 |
| Length | 118 |
| Max. P | 0.939117 |

| Location | 22,201,887 – 22,202,005 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.37 |
| Mean single sequence MFE | -33.55 |
| Consensus MFE | -23.30 |
| Energy contribution | -24.42 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.873707 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
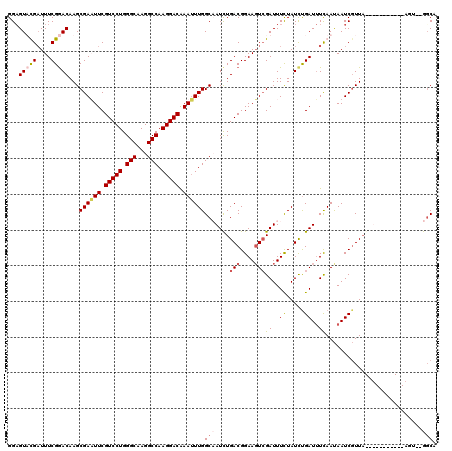
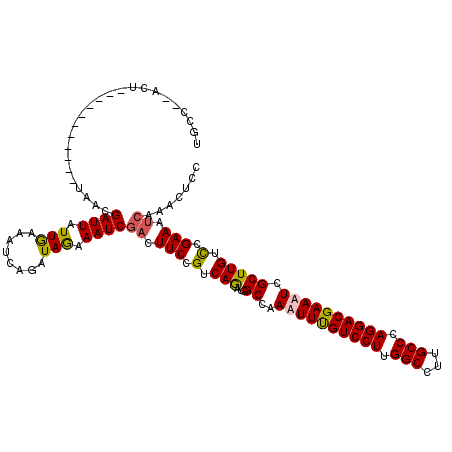
>3L_DroMel_CAF1 22201887 118 + 23771897 GGAGUUUGAUUUCGGACAAGCGAAUUCGUCCUGGGCAAGGCCAAGGACAAAUUUGGCCAUUUGACGGAAUUCGAUUUUUAUUUGAUUUCAAUAAUCGUUAGUUAAAAUCUUAGU--GGCA ...(((((....)))))...((((((.(((((.(((...))).))))).))))))((((((((((......(((((.....(((....))).)))))...))))).......))--))). ( -31.61) >DroSec_CAF1 60819 120 + 1 GGAGUUUGAUUUCGGACAAGCGAAUUCGUCCUGGGCAAGGCCAAGGACAAAUUUGGCCAUCUGACGGAAGUCGAUUUCUAUCCGAUUUCAAUAAUCGUUAGUUACAAUCUUAGAUAGGCA ...(((((....)))))...((((((.(((((.(((...))).))))).))))))((((((((((....)))((((.(((..(((((.....))))).)))....))))..)))).))). ( -39.20) >DroEre_CAF1 60620 104 + 1 GGAGUACGUUUUCGGACAAGCGAUUUCGUCCUGGGCAAGGCCAAGGACAAAGUUGGCAAUCUGACGGAAGUGGAUUUCUAUCUGAUUUUAAUAAUCGUUA----------------GUCA ((((.....)))).(((.(((((((..(((((.(((...))).)))))..((.(((.(((((.((....))))))).))).)).........))))))).----------------))). ( -31.90) >DroYak_CAF1 72348 107 + 1 GGCGUACGAUUUCGAACAAGCGAUUUCGUCCUGGGCAAGGCCAAGGACGAAUUUGGCAAUCUGACGGAAGUCGAUUUCUAUCUGAUAUCAACAAUCUUUU-----------ACU--GGCA .((((.((....)).))...(((.((((((((.(((...))).)))))))).))))).....(((....)))(((.((.....)).)))...........-----------...--.... ( -31.50) >consensus GGAGUACGAUUUCGGACAAGCGAAUUCGUCCUGGGCAAGGCCAAGGACAAAUUUGGCAAUCUGACGGAAGUCGAUUUCUAUCUGAUUUCAAUAAUCGUUA___________AGU__GGCA ...(((((....)))))...((((((.(((((.(((...))).))))).))))))(((....(((....)))((..((.....))..))...........................))). (-23.30 = -24.42 + 1.13)


| Location | 22,201,887 – 22,202,005 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.37 |
| Mean single sequence MFE | -28.95 |
| Consensus MFE | -19.41 |
| Energy contribution | -19.48 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.67 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939117 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
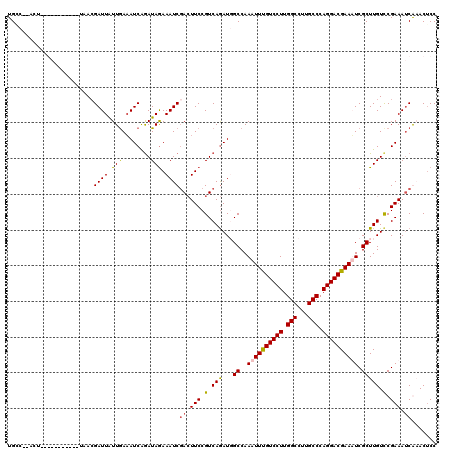
>3L_DroMel_CAF1 22201887 118 - 23771897 UGCC--ACUAAGAUUUUAACUAACGAUUAUUGAAAUCAAAUAAAAAUCGAAUUCCGUCAAAUGGCCAAAUUUGUCCUUGGCCUUGCCCAGGACGAAUUCGCUUGUCCGAAAUCAAACUCC ....--.....(((((((((....)....))))))))...........((.(((.(.((...(((..((((((((((.(((...))).)))))))))).))))).).))).))....... ( -26.30) >DroSec_CAF1 60819 120 - 1 UGCCUAUCUAAGAUUGUAACUAACGAUUAUUGAAAUCGGAUAGAAAUCGACUUCCGUCAGAUGGCCAAAUUUGUCCUUGGCCUUGCCCAGGACGAAUUCGCUUGUCCGAAAUCAAACUCC ...........((((((.....)))))).((((..(((((((...((((((....))).)))(((..((((((((((.(((...))).)))))))))).))))))))))..))))..... ( -35.50) >DroEre_CAF1 60620 104 - 1 UGAC----------------UAACGAUUAUUAAAAUCAGAUAGAAAUCCACUUCCGUCAGAUUGCCAACUUUGUCCUUGGCCUUGCCCAGGACGAAAUCGCUUGUCCGAAAACGUACUCC .(((----------------...(((((.....((((.((..(((......)))..)).)))).......(((((((.(((...))).))))))))))))...))).............. ( -25.00) >DroYak_CAF1 72348 107 - 1 UGCC--AGU-----------AAAAGAUUGUUGAUAUCAGAUAGAAAUCGACUUCCGUCAGAUUGCCAAAUUCGUCCUUGGCCUUGCCCAGGACGAAAUCGCUUGUUCGAAAUCGUACGCC ....--.((-----------(...((((((((((.((.....)).)))))).....((..((.((....((((((((.(((...))).))))))))...))..))..)))))).)))... ( -29.00) >consensus UGCC__ACU___________UAACGAUUAUUGAAAUCAGAUAGAAAUCGACUUCCGUCAGAUGGCCAAAUUUGUCCUUGGCCUUGCCCAGGACGAAAUCGCUUGUCCGAAAUCAAACUCC ........................((((.(((........))).))))((.(((.(.(((...((..((((((((((.(((...))).)))))))))).))))).).))).))....... (-19.41 = -19.48 + 0.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:15:32 2006