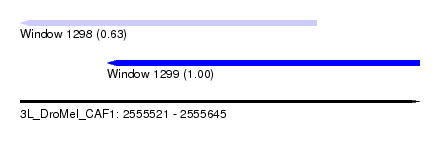
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 2,555,521 – 2,555,645 |
| Length | 124 |
| Max. P | 0.999833 |

| Location | 2,555,521 – 2,555,613 |
|---|---|
| Length | 92 |
| Sequences | 6 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 68.18 |
| Mean single sequence MFE | -13.05 |
| Consensus MFE | -6.91 |
| Energy contribution | -7.52 |
| Covariance contribution | 0.61 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.629802 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
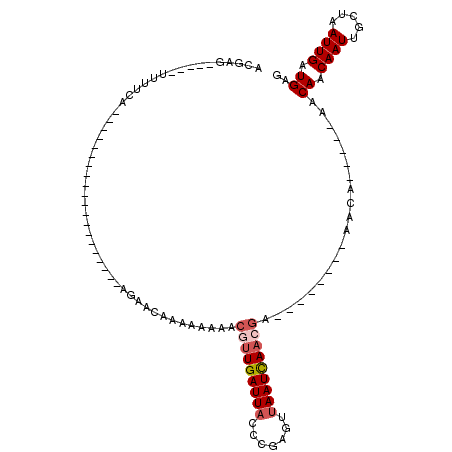
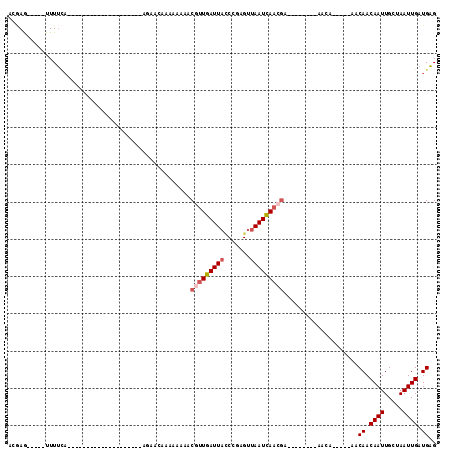
>3L_DroMel_CAF1 2555521 92 - 23771897 ACGAG-----UUUUCA--GUUUUUUUUCAAAAA---AAAAAAAAAAAAACGUUGAUUACCCAAGUUAAUCAACGA--------AACU-----AACAACAAUUGCUAAUUGAUGAG ...((-----((((..--(((((((((......---.....)))))))))((((((((.......))))))))))--------))))-----..((.((((.....)))).)).. ( -15.90) >DroVir_CAF1 22242 110 - 1 UAAAGUGCAGUUUGAAAA----UAAAACAGAAAACAAAAAAAAAAACUACUUUGAUUU-CUGGGUUAAUUAACUAACAGCAGAAGCAGCAGCAGCAACAAUUGGUAAUUGAUGCG .....(((((((......----(((..((((((.((((............)))).)))-)))..)))...))))....((....)).)))...(((.((((.....)))).))). ( -19.00) >DroSec_CAF1 17480 75 - 1 ACGAA-----UUUUCA----------------------AACAACAAAAACGUUGAUUACCCAAGUUAAUCAACGA--------AACU-----AACAACAAUUGCUAAUUGAUGAG .....-----......----------------------...........(((((((((.......))))))))).--------....-----..((.((((.....)))).)).. ( -10.60) >DroEre_CAF1 31500 77 - 1 ACGAG-----UUUUCA--------------------AGCACAACAAAAACGUUGAUUACCCGAGUUAAUCAACGA--------AACA-----AACAACAAUUGCUAAUUGAUGAG .....-----......--------------------.............(((((((((.......))))))))).--------....-----..((.((((.....)))).)).. ( -10.60) >DroYak_CAF1 15898 77 - 1 ACGAG-----UUUUCA--------------------AGCACAACAAAAACGUUGAUUACCCGGGUUAAUCAACGA--------AACA-----UACAACAAUUGCUAAUUGAUGAG .....-----......--------------------.............((((((((((....).))))))))).--------....-----..((.((((.....)))).)).. ( -12.00) >DroAna_CAF1 16390 73 - 1 ---AG-----UUUUUC--------------------AGCA-AAAAAACGCUUUGAUUAUCCGGGUUAAUUAUCGA--------AACU-----AACAACAAUUGGUAAUUGAUGAG ---.(-----(((((.--------------------....-.))))))...............((((((((((((--------....-----........))))))))))))... ( -10.20) >consensus ACGAG_____UUUUCA____________________AGAACAAAAAAAACGUUGAUUACCCGAGUUAAUCAACGA________AACA_____AACAACAAUUGCUAAUUGAUGAG .................................................(((((((((.......)))))))))....................((.((((.....)))).)).. ( -6.91 = -7.52 + 0.61)


| Location | 2,555,548 – 2,555,645 |
|---|---|
| Length | 97 |
| Sequences | 3 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 73.63 |
| Mean single sequence MFE | -15.87 |
| Consensus MFE | -12.00 |
| Energy contribution | -12.33 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.76 |
| SVM decision value | 4.20 |
| SVM RNA-class probability | 0.999833 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
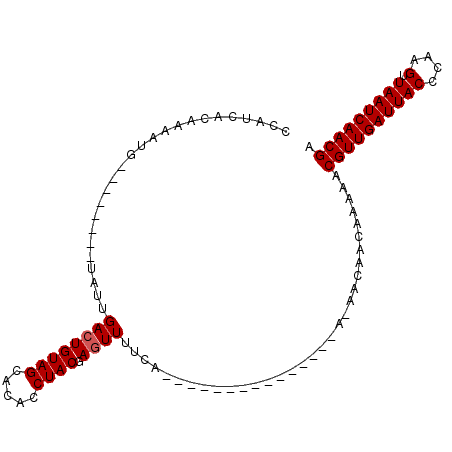
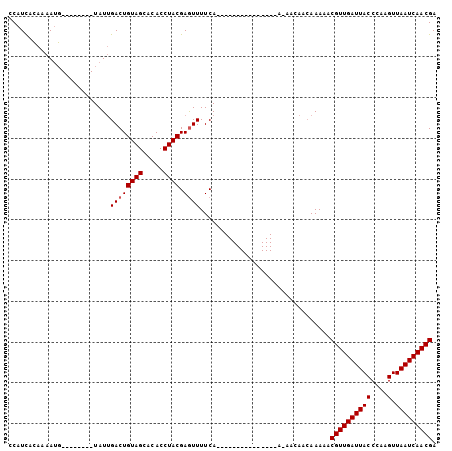
>3L_DroMel_CAF1 2555548 97 - 23771897 CCAUCUCAAAAUG--------UAUUGACUGUAGCACACCUACGAGUUUUCAGUUUUUUUUCAAAAAAAAAAAAAAAAAACGUUGAUUACCCAAGUUAAUCAACGA ...........((--------....((((((((.....)))).))))..))(((((((((...........)))))))))((((((((.......)))))))).. ( -16.20) >DroSec_CAF1 17507 80 - 1 CCAUUACAAAACG--------UAUUGACUGUAGCACACCUACGAAUUUUCA-----------------AACAACAAAAACGUUGAUUACCCAAGUUAAUCAACGA .((.(((.....)--------)).))..(((((.....)))))........-----------------...........(((((((((.......))))))))). ( -13.40) >DroYak_CAF1 15925 90 - 1 CCAUCAACAAAUUGGGUAGAUUAUAGACUGUAGCACACCUACGAGUUUUCA---------------AGCACAACAAAAACGUUGAUUACCCGGGUUAAUCAACGA (((.........)))((.(.....(((((((((.....)))).)))))...---------------..)))........((((((((((....).))))))))). ( -18.00) >consensus CCAUCACAAAAUG________UAUUGACUGUAGCACACCUACGAGUUUUCA_______________A_AACAACAAAAACGUUGAUUACCCAAGUUAAUCAACGA .........................((((((((.....)))).))))................................((((((((((....).))))))))). (-12.00 = -12.33 + 0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:52:35 2006