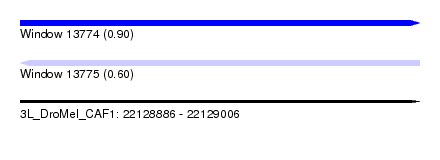

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 22,128,886 – 22,129,006 |
| Length | 120 |
| Max. P | 0.900290 |

| Location | 22,128,886 – 22,129,006 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.98 |
| Mean single sequence MFE | -29.77 |
| Consensus MFE | -25.82 |
| Energy contribution | -25.68 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.900290 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
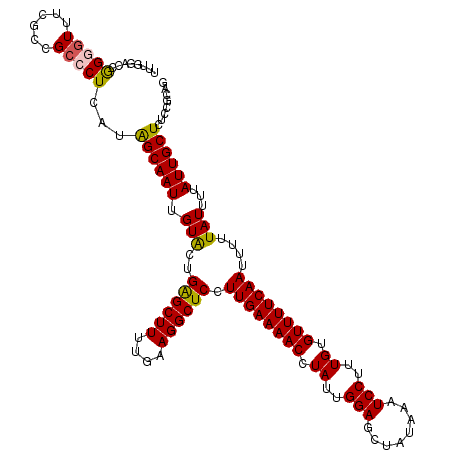
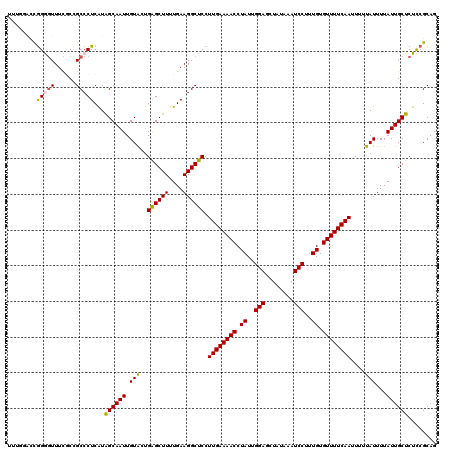
>3L_DroMel_CAF1 22128886 120 + 23771897 UUUGGACCGGGGUUUCACCGCCCUCAUAGCAAUUGUACUGAGCUUUUGAAGGCUCCUUGAAAACCUAUUGGAGCUAUAAAUCCUUUGUGUUUUCAAUUUUUAUUUUAUUGCUCUCCGCAG ...(((..(((((......)))))...((((((.(((..((((((....)))))).((((((((.((..(((........)))..)).))))))))....)))...)))))).))).... ( -31.30) >DroSec_CAF1 114695 120 + 1 UUUGGACCGGGGUUUCGCCGCCCUCAUGGCAAUUGUACUGAGCUUUUGAAGGCUCCUUGAAAACCUAUUGGAGCUAUAAAUCCUUUGUGUUUUCAAUUUUUAUUUUAUUGCUCUCCGCAG ...(((..(((((......)))))...((((((.(((..((((((....)))))).((((((((.((..(((........)))..)).))))))))....)))...)))))).))).... ( -31.30) >DroSim_CAF1 122794 120 + 1 UUUGGACCGGGGUUUCGCCGCCCUCAUGGCAAUUGUACUGAGCUUUUGAAGGCUCCUUGAAAACCUAUUGGAGCUAUAAAUCCUUUGUGUUUUCAAUUUUUAUUUUAUUGCUCUCCGCAG ...(((..(((((......)))))...((((((.(((..((((((....)))))).((((((((.((..(((........)))..)).))))))))....)))...)))))).))).... ( -31.30) >DroEre_CAF1 113588 115 + 1 -----GUUAGGGUUUCGCCGCCCUCAUAGCAAUUGUACUGAGCUUUUGAAGGCUCCUUGAAAACCUAUUGGAGCUAUAAAUCCUUUGUGUUUUCAAUUUUUAUUUUAUUGCUCUCCGCAG -----((.(((((......))))).))((((((.(((..((((((....)))))).((((((((.((..(((........)))..)).))))))))....)))...))))))........ ( -28.40) >DroYak_CAF1 162317 120 + 1 UUGAAACUGGAGUUUCGCCGCCCUCAUAGCAAUUGUACUGAGCUUUUGAAGGCUCCUUGAAAACCUAUUGGAGCUAUAAAUCCUUUGUGUUUUCAAUUUUUAUUUUAUUGCUCUCCGCAG ..(((((....)))))...((......((((((.(((..((((((....)))))).((((((((.((..(((........)))..)).))))))))....)))...))))))....)).. ( -28.20) >DroAna_CAF1 132051 114 + 1 ------UCGGAUUUUUCCAGCCCUCGUAGCAAUUGUGCCGGGCUUUUGAAGGCUCCUUGAAAACCUAUUGGAGCUAUAAAUCCUUUGUGUUUUCAAUUUUUAUUUUAUUGCUCUUCGCAG ------..(((.(((((.(((((..(((.(....)))).)))))...))))).)))((((((((.((..(((........)))..)).))))))))............(((.....))). ( -28.10) >consensus UUUGGACCGGGGUUUCGCCGCCCUCAUAGCAAUUGUACUGAGCUUUUGAAGGCUCCUUGAAAACCUAUUGGAGCUAUAAAUCCUUUGUGUUUUCAAUUUUUAUUUUAUUGCUCUCCGCAG ........(((((......)))))...((((((.(((..((((((....)))))).((((((((.((..(((........)))..)).))))))))....)))...))))))........ (-25.82 = -25.68 + -0.14)

| Location | 22,128,886 – 22,129,006 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.98 |
| Mean single sequence MFE | -28.83 |
| Consensus MFE | -23.22 |
| Energy contribution | -23.47 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.21 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.597431 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
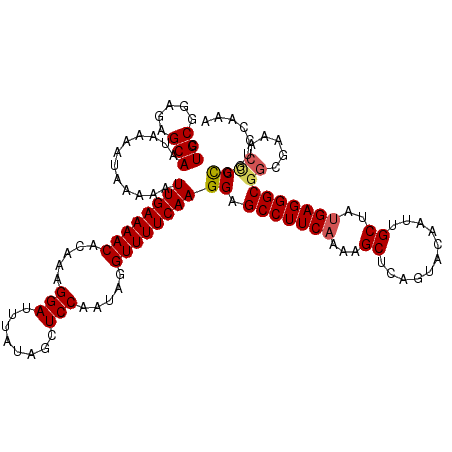
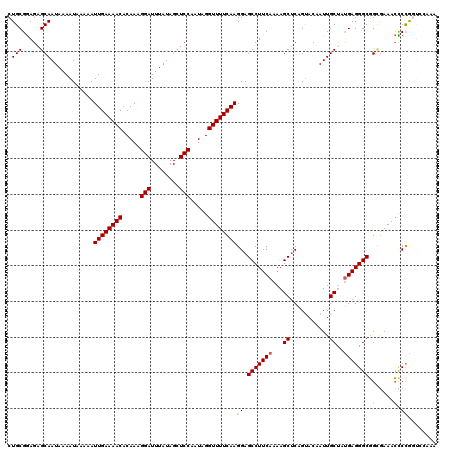
>3L_DroMel_CAF1 22128886 120 - 23771897 CUGCGGAGAGCAAUAAAAUAAAAAUUGAAAACACAAAGGAUUUAUAGCUCCAAUAGGUUUUCAAGGAGCCUUCAAAAGCUCAGUACAAUUGCUAUGAGGGCGGUGAAACCCCGGUCCAAA ....(((.((((((..........((((((((.....(((........))).....)))))))).((((........))))......))))))....(((.(......))))..)))... ( -29.60) >DroSec_CAF1 114695 120 - 1 CUGCGGAGAGCAAUAAAAUAAAAAUUGAAAACACAAAGGAUUUAUAGCUCCAAUAGGUUUUCAAGGAGCCUUCAAAAGCUCAGUACAAUUGCCAUGAGGGCGGCGAAACCCCGGUCCAAA ....((((.(((((..........((((((((.....(((........))).....)))))))).((((........))))......))))))....(((.(......))))..)))... ( -28.30) >DroSim_CAF1 122794 120 - 1 CUGCGGAGAGCAAUAAAAUAAAAAUUGAAAACACAAAGGAUUUAUAGCUCCAAUAGGUUUUCAAGGAGCCUUCAAAAGCUCAGUACAAUUGCCAUGAGGGCGGCGAAACCCCGGUCCAAA ....((((.(((((..........((((((((.....(((........))).....)))))))).((((........))))......))))))....(((.(......))))..)))... ( -28.30) >DroEre_CAF1 113588 115 - 1 CUGCGGAGAGCAAUAAAAUAAAAAUUGAAAACACAAAGGAUUUAUAGCUCCAAUAGGUUUUCAAGGAGCCUUCAAAAGCUCAGUACAAUUGCUAUGAGGGCGGCGAAACCCUAAC----- ....((...((.............((((((((.....(((........))).....))))))))...(((((((..(((...........))).))))))).))....)).....----- ( -26.00) >DroYak_CAF1 162317 120 - 1 CUGCGGAGAGCAAUAAAAUAAAAAUUGAAAACACAAAGGAUUUAUAGCUCCAAUAGGUUUUCAAGGAGCCUUCAAAAGCUCAGUACAAUUGCUAUGAGGGCGGCGAAACUCCAGUUUCAA ....((((.((.............((((((((.....(((........))).....))))))))...(((((((..(((...........))).))))))).))....))))........ ( -30.80) >DroAna_CAF1 132051 114 - 1 CUGCGAAGAGCAAUAAAAUAAAAAUUGAAAACACAAAGGAUUUAUAGCUCCAAUAGGUUUUCAAGGAGCCUUCAAAAGCCCGGCACAAUUGCUACGAGGGCUGGAAAAAUCCGA------ .(((.....)))............((((((((.....(((........))).....))))))))(((...(((...(((((((((....))))....))))).)))...)))..------ ( -30.00) >consensus CUGCGGAGAGCAAUAAAAUAAAAAUUGAAAACACAAAGGAUUUAUAGCUCCAAUAGGUUUUCAAGGAGCCUUCAAAAGCUCAGUACAAUUGCUAUGAGGGCGGCGAAACCCCGGUCCAAA .(((.....)))............((((((((.....(((........))).....))))))))((.(((((((...((...........))..)))))))((.....))))........ (-23.22 = -23.47 + 0.25)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:14:59 2006