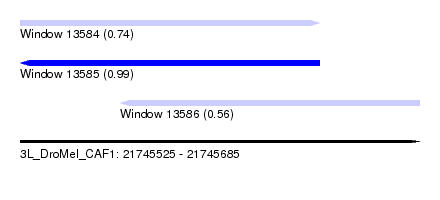
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,745,525 – 21,745,685 |
| Length | 160 |
| Max. P | 0.988676 |

| Location | 21,745,525 – 21,745,645 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 99.00 |
| Mean single sequence MFE | -37.94 |
| Consensus MFE | -35.92 |
| Energy contribution | -36.32 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.744218 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21745525 120 + 23771897 CGAAAAUUGUCACGGUUUCGCGGUCGAUGGCUCGCUUGCUGUGAAGGUGCAAAUUUAAUGUUUGCCAAUUUAGCAGCGCUCUGCUGUCACCUUUGGUGUCCAUUAGCGAGCGUUUAUAUG ...(((((((((((((...(((((.....)).)))..)))))))....((((((.....))))))))))))...(((((((.((((.((((...)))).....)))))))))))...... ( -39.20) >DroSec_CAF1 29897 120 + 1 CGAAAAUUGUCACGGUUUCGCGGUCGAUGGCUCGCUCGCUGUGAAGGUGCAAAUUUAAUGUUUGCCAAUUUAGCAGCUCUCUGCUGUCACCUUUGGUGUCCAUUAGCGAGCGUUUAUAUG ........((((((..(....)..)).)))).(((((((((.((((((((((((.....)))))).......(((((.....))))).))))))((...))..)))))))))........ ( -36.80) >DroSim_CAF1 32145 120 + 1 CGAAAAUUGUCACGGUUUCGCGGUCGAUGGCUCGCUCGCUGUGAAGGUGCAAAUUUAAUGUUUGCCAAUUUAGCAGCGCUCUGCUGUCACCUUUGGUGUCCAUUAGCGAGCGUUUAUAUG ...(((((((((((((...(((((.....)).)))..)))))))....((((((.....))))))))))))...(((((((.((((.((((...)))).....)))))))))))...... ( -38.50) >DroEre_CAF1 13603 120 + 1 CGAAAAUUGUCACGGUUUCGCGGUCGAUGGCUCGCUCGCUGUGAAGGUGCAAAUUUAAUGUUUGCCAAUUUAGCAGCGCUCUGCUGUCACCUUUGGUGUCCAUUAGCGAGCGUUUAUAUG ...(((((((((((((...(((((.....)).)))..)))))))....((((((.....))))))))))))...(((((((.((((.((((...)))).....)))))))))))...... ( -38.50) >DroYak_CAF1 90597 120 + 1 CGAAAAUUGUCACGGUUUCGCGUUCGAUGGCUCGCUCGCUGUGAAGGUGCAAAUUUAAUGUUUGCCAAUUUAGCAGCGCUCUGCUGUCACCUUUGGUGUCCAUUAGCGAGCGUUUAUAUG ........(((((((........))).)))).(((((((((.((((((((((((.....)))))).....((((((....))))))..))))))((...))..)))))))))........ ( -36.70) >consensus CGAAAAUUGUCACGGUUUCGCGGUCGAUGGCUCGCUCGCUGUGAAGGUGCAAAUUUAAUGUUUGCCAAUUUAGCAGCGCUCUGCUGUCACCUUUGGUGUCCAUUAGCGAGCGUUUAUAUG ...(((((((((((((...(((((.....)).)))..)))))))....((((((.....))))))))))))...(((((((.((((.((((...)))).....)))))))))))...... (-35.92 = -36.32 + 0.40)

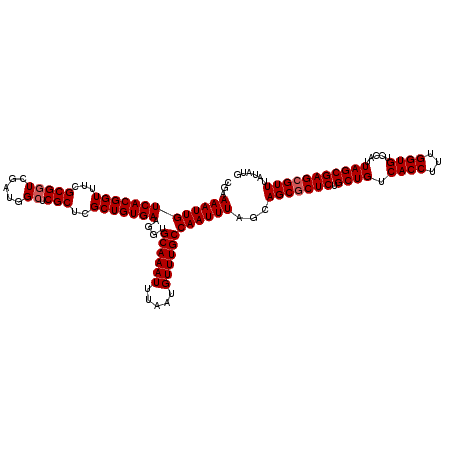

| Location | 21,745,525 – 21,745,645 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 99.00 |
| Mean single sequence MFE | -32.78 |
| Consensus MFE | -31.88 |
| Energy contribution | -32.08 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.97 |
| SVM decision value | 2.13 |
| SVM RNA-class probability | 0.988676 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21745525 120 - 23771897 CAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCACAGCAAGCGAGCCAUCGACCGCGAAACCGUGACAAUUUUCG .........(((((((((((....((((.(....)(((((....)))))...))))....)))))...((((.........)))))))))).......((((....)))).......... ( -31.70) >DroSec_CAF1 29897 120 - 1 CAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGAGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCACAGCGAGCGAGCCAUCGACCGCGAAACCGUGACAAUUUUCG ........((((((((..(((...)))(((((...(((((....)))))......(((((.......))))))))))...))))))))..........((((....)))).......... ( -33.00) >DroSim_CAF1 32145 120 - 1 CAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCACAGCGAGCGAGCCAUCGACCGCGAAACCGUGACAAUUUUCG ........((((((((..(((...)))(((((...(((((....)))))......(((((.......))))))))))...))))))))..........((((....)))).......... ( -33.00) >DroEre_CAF1 13603 120 - 1 CAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCACAGCGAGCGAGCCAUCGACCGCGAAACCGUGACAAUUUUCG ........((((((((..(((...)))(((((...(((((....)))))......(((((.......))))))))))...))))))))..........((((....)))).......... ( -33.00) >DroYak_CAF1 90597 120 - 1 CAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCACAGCGAGCGAGCCAUCGAACGCGAAACCGUGACAAUUUUCG ........((((((((..(((...)))(((((...(((((....)))))......(((((.......))))))))))...))))))))..........((((....)))).......... ( -33.20) >consensus CAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCACAGCGAGCGAGCCAUCGACCGCGAAACCGUGACAAUUUUCG ........((((((((..(((...)))(((((...(((((....)))))......(((((.......))))))))))...))))))))..........((((....)))).......... (-31.88 = -32.08 + 0.20)

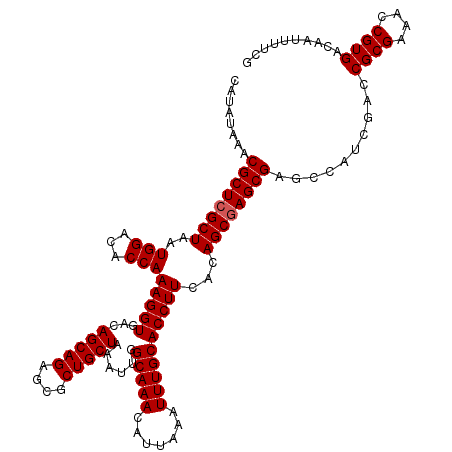

| Location | 21,745,565 – 21,745,685 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.92 |
| Mean single sequence MFE | -32.94 |
| Consensus MFE | -31.42 |
| Energy contribution | -31.70 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.563241 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21745565 120 - 23771897 UGCCAGAUCCGGAGCUGGCCUGCCAAUGCUCGGCACUUUGCAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCAC .(((((........))))).(((((((...((((.((((((.................(((...)))..(....).)))))).))))....)))))))...................... ( -34.50) >DroSec_CAF1 29937 120 - 1 UGCCAGAUCCGGAGCUGGCCUGCCAAUGCUCGGCACUUUGCAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGAGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCAC .(((((........))))).(((((((...((((.((((((.................(((...)))..(....).)))))).))))....)))))))...................... ( -33.70) >DroSim_CAF1 32185 120 - 1 UGCCAGAUCCGGAGCUGGCCUGCCAAUGCUCGGCACUUUGCAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCAC .(((((........))))).(((((((...((((.((((((.................(((...)))..(....).)))))).))))....)))))))...................... ( -34.50) >DroEre_CAF1 13643 120 - 1 UGGCAGAUCCGAAGCUGGCCUGCCAAUGCUCGGCACUUUGCAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCAC ((((((..((((.(((((....)))..))))))...............((((((((..(((...)))..(....)))).))))))))))).....(((((.......)))))........ ( -33.30) >DroYak_CAF1 90637 120 - 1 UUUCAGAUCCCAAGCUGGCCUGCCAAUGCUCGGCACUUUGCAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCAC ..((((........))))..(((((((...((((.((((((.................(((...)))..(....).)))))).))))....)))))))...................... ( -28.70) >consensus UGCCAGAUCCGGAGCUGGCCUGCCAAUGCUCGGCACUUUGCAUAUAAACGCUCGCUAAUGGACACCAAAGGUGACAGCAGAGCGCUGCUAAAUUGGCAAACAUUAAAUUUGCACCUUCAC .(((((........))))).(((((((...((((.((((((.................(((...)))..(....).)))))).))))....)))))))...................... (-31.42 = -31.70 + 0.28)


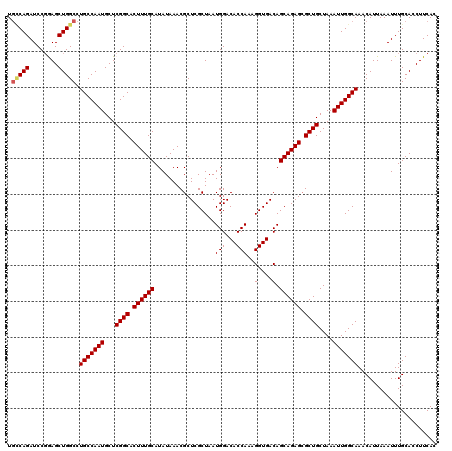
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:12:00 2006