
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,741,162 – 21,741,322 |
| Length | 160 |
| Max. P | 0.899008 |

| Location | 21,741,162 – 21,741,282 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.74 |
| Mean single sequence MFE | -31.22 |
| Consensus MFE | -21.42 |
| Energy contribution | -23.42 |
| Covariance contribution | 2.00 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.82 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.899008 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21741162 120 - 23771897 CCACUAAAGUGAUCCCCGGGCAUUUGACCAGUCGUUGUCUGUCCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUCAGGCUCAGGCUCAGGCUCAGACUCCCGGCUUCUGUCACUU ......(((((((..(((((..(((((((((((...(((((..........................)))))...))))..(((....)))..)).)))))..))))).....))))))) ( -33.27) >DroSec_CAF1 23072 114 - 1 CCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUCAGGCUC------AGGCUCAGACUCCCGGCUUCUGUCACUU ......(((((((..(((((..((((((((((((..(((((.((....................)).)))))..))))).......------.)).)))))..))))).....))))))) ( -34.75) >DroSim_CAF1 27949 114 - 1 CCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUCAGGCUC------AGGCUCAGACUCCCGGCUUCUGUCACUU ......(((((((..(((((..((((((((((((..(((((.((....................)).)))))..))))).......------.)).)))))..))))).....))))))) ( -34.75) >DroEre_CAF1 9197 101 - 1 CCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCGCAUCUCAUCUCAUCUCAG-------CUC------------AGACUCAGGCUCCCGGCUUCUGUCACUU ......(((((((..(((((.........(((((..(((((.((.........................)-------).)------------))))..)))))))))).....))))))) ( -26.51) >DroYak_CAF1 81108 108 - 1 CCACUCAAGGGAUCCCCGGGCAUUUGACCAGUCUUUGUCCGUGCCUCCAUCUCAGCUCAUCUCAGCUCAGACUCAGACUC------------AGACUCAGGCUCUCGGCUUCUGUCACUU .........(((..(.((((((...((....))..)))))).)..))).(((.((((......)))).))).........------------.(((..((((.....))))..))).... ( -26.80) >consensus CCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUCAGGCUC______AGGCUCAGACUCCCGGCUUCUGUCACUU ......(((((((..(((((..((((((((((((..(((((.((....................)).)))))..)))))..............)).)))))..))))).....))))))) (-21.42 = -23.42 + 2.00)
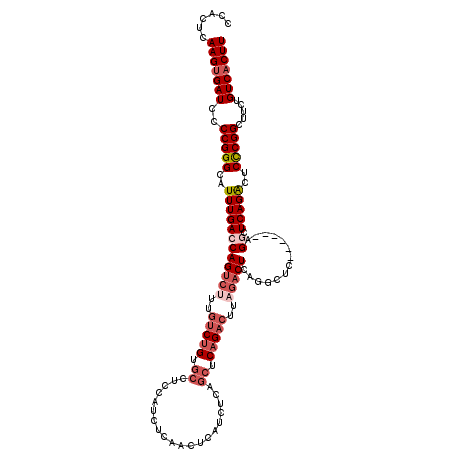


| Location | 21,741,202 – 21,741,322 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.83 |
| Mean single sequence MFE | -33.42 |
| Consensus MFE | -26.50 |
| Energy contribution | -27.34 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.758742 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21741202 120 - 23771897 AUAAGCAUGAUCAGAUGACUGCCUUCGUGCGCCGAGAUGCCCACUAAAGUGAUCCCCGGGCAUUUGACCAGUCGUUGUCUGUCCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUC ....((((((.(((....)))...))))))((.((((((..(((....)))......(((((..((((.....))))..))))).............)))))).)).............. ( -29.50) >DroSec_CAF1 23106 120 - 1 AUAAGCAUGAUCAGAUGACUGCCUUCGUGCGCCGAGAUGCCCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUC ....((((((.(((....)))...))))))....((((((((...............))))))))....(((((..(((((.((....................)).)))))..))))). ( -34.51) >DroSim_CAF1 27983 120 - 1 AUAAGCAUGAUCAGAUGACUGCCUUCGUGCGCCGAGAUGCCCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUC ....((((((.(((....)))...))))))....((((((((...............))))))))....(((((..(((((.((....................)).)))))..))))). ( -34.51) >DroEre_CAF1 9225 113 - 1 AUAAGCAUGAUCAGAUGACUGCCUUCGUGUGCCGAGAUGCCCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCGCAUCUCAUCUCAUCUCAG-------CUC ...(((.(((...(((((..(((.....).)).(((((((.(((....)))......((((((..(((........))).)))))).....)))))))...)))))))))-------)). ( -31.90) >DroYak_CAF1 81136 117 - 1 AUGAGCAUGAUCAGAUG---GCCUUCGUGCGCCGAGAUGCCCACUCAAGGGAUCCCCGGGCAUUUGACCAGUCUUUGUCCGUGCCUCCAUCUCAGCUCAUCUCAGCUCAGACUCAGACUC .(((((.(((...((((---((......))((.(((((((((......))).....((((((...((....))..))))))......)))))).)).))))))))))))........... ( -36.70) >consensus AUAAGCAUGAUCAGAUGACUGCCUUCGUGCGCCGAGAUGCCCACUCAAGUGAUCCCCGGGCAUUUGACCAGUCUUUGUCUGUGCCUCCAUCUCAACUCAUCUCAGCUCAGACUUAGACUC ....((((((.(((....)))...))))))((.((((((..(((....)))......((((((..(((........))).))))))...........)))))).)).............. (-26.50 = -27.34 + 0.84)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:11:44 2006