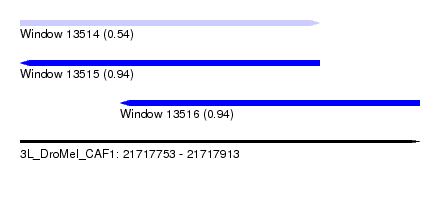
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,717,753 – 21,717,913 |
| Length | 160 |
| Max. P | 0.940283 |

| Location | 21,717,753 – 21,717,873 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.93 |
| Mean single sequence MFE | -18.92 |
| Consensus MFE | -10.82 |
| Energy contribution | -10.83 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.535085 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21717753 120 + 23771897 CAGCCCAGUCCACCCACUCCUACUCCUACUGCAGAGGCUGUCAUUGAAUUUUAAUUUAAUCCAAUGCAAGUCGAAUGCAAACAAAAAAUUGCAUCUGACUCUAUUACAAUGCAAAGACAA .((((((((..................))))....))))((((((((((....)))))))....(((((((((.((((((........)))))).))))).........))))..))).. ( -22.57) >DroPse_CAF1 26114 108 + 1 ------GCUCCUCCUUCUCCUGCU------UCUGCUGCUGUCAUUGAAUUUUAAUUUAAUUCAAUUCAAGCCAAAAACAAACACAAAAUUGCAUCUGACUCUAUUACAAUGGCAAGACAA ------...............((.------...))...((((((((((((.......))))))))....((((....................................))))..)))). ( -12.72) >DroSim_CAF1 3963 120 + 1 CAGCCCAGUCCACCCACUCCUACUCCUACUGCAGAGGCUGUCAUUGAAUUUUAAUUUAAUCCAAUGCAAGCCAAAUGCAAACAAAAAAUUGCAUCUGACUCUAUUACAAUGCAAAGACAA .......(((...................((((..((((..(((((.(((.......))).)))))..))))...)))).........((((((.((.........)))))))).))).. ( -22.00) >DroYak_CAF1 50420 108 + 1 ------------ACUACUUCCACUCCUACUCCAGAGGCUGUCAUUGAAUUUUAAUUUAAUCCAAUGCAAGCCAAAUGCAAACAAAAAAUUGCAUCUGACUCUAUUACAAUGCAAAGACGA ------------...................((((((((..(((((.(((.......))).)))))..))))...(((((........)))))))))....................... ( -18.40) >consensus ______AGUCCACCCACUCCUACUCCUACUGCAGAGGCUGUCAUUGAAUUUUAAUUUAAUCCAAUGCAAGCCAAAUGCAAACAAAAAAUUGCAUCUGACUCUAUUACAAUGCAAAGACAA ......................................(((((((((((....)))))))....((((((.((.((((((........)))))).)).)).........))))..)))). (-10.82 = -10.83 + 0.00)

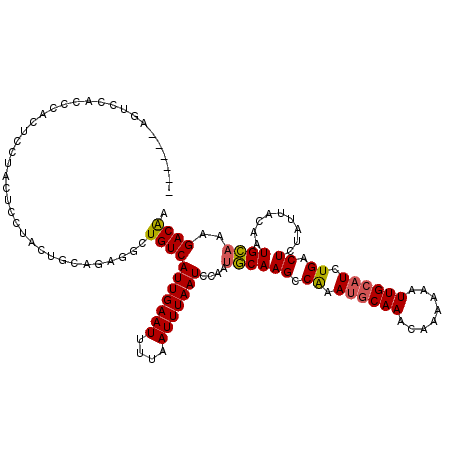
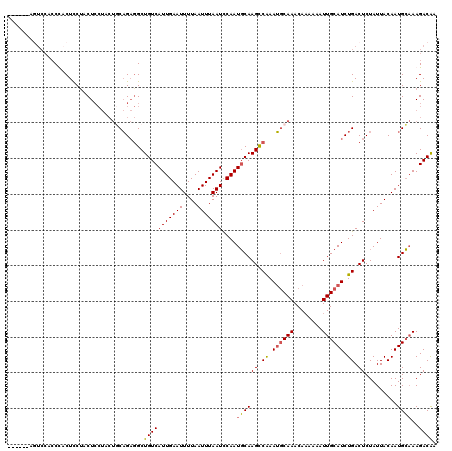
| Location | 21,717,753 – 21,717,873 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.93 |
| Mean single sequence MFE | -31.32 |
| Consensus MFE | -22.61 |
| Energy contribution | -22.55 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.940283 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21717753 120 - 23771897 UUGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUCGACUUGCAUUGGAUUAAAUUAAAAUUCAAUGACAGCCUCUGCAGUAGGAGUAGGAGUGGGUGGACUGGGCUG ..((((.(.((((((....(((((..((((((........))))))..))))).(((((((((.......)))))))))))..(((((.(....).)).))))))).).))))....... ( -32.40) >DroPse_CAF1 26114 108 - 1 UUGUCUUGCCAUUGUAAUAGAGUCAGAUGCAAUUUUGUGUUUGUUUUUGGCUUGAAUUGAAUUAAAUUAAAAUUCAAUGACAGCAGCAGA------AGCAGGAGAAGGAGGAGC------ ((.((((((..((((..(((((.((((((((....)))))))).)))))(((...((((((((.......))))))))...))).)))).------.)))))).))........------ ( -27.70) >DroSim_CAF1 3963 120 - 1 UUGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUUGGCUUGCAUUGGAUUAAAUUAAAAUUCAAUGACAGCCUCUGCAGUAGGAGUAGGAGUGGGUGGACUGGGCUG ..((((.(.((((((....(((((((((((((........))))))))))))).(((((((((.......)))))))))))..(((((.(....).)).))))))).).))))....... ( -35.40) >DroYak_CAF1 50420 108 - 1 UCGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUUGGCUUGCAUUGGAUUAAAUUAAAAUUCAAUGACAGCCUCUGGAGUAGGAGUGGAAGUAGU------------ ......((((.((((....(((((((((((((........))))))))))))).(((((((((.......)))))))))))))(((((......)))).)...)))).------------ ( -29.80) >consensus UUGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUUGGCUUGCAUUGGAUUAAAUUAAAAUUCAAUGACAGCCUCUGCAGUAGGAGUAGGAGUAGGUGGACU______ ..(((..............(((((((((((((........)))))))))))))..((((((((.......)))))))))))....................................... (-22.61 = -22.55 + -0.06)



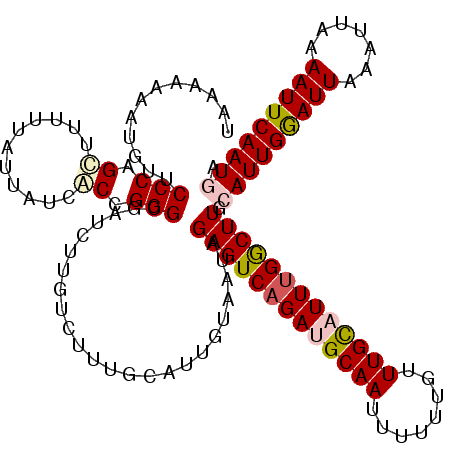
| Location | 21,717,793 – 21,717,913 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.67 |
| Mean single sequence MFE | -26.78 |
| Consensus MFE | -22.44 |
| Energy contribution | -23.04 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938761 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21717793 120 - 23771897 UAAAAAAAUGUUCCCAGCUUUUUAUUAUCACCGGGCGAUCUUGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUCGACUUGCAUUGGAUUAAAUUAAAAUUCAAUGA ((((((..((....))..)))))).(((.((...((((........))))...)).)))(((((..((((((........))))))..))))).(((((((((.......))))))))). ( -26.10) >DroPse_CAF1 26142 120 - 1 UACCAAAAUUUUCCCAGCUUUUUAUUAUCGCCGGGUGAUCUUGUCUUGCCAUUGUAAUAGAGUCAGAUGCAAUUUUGUGUUUGUUUUUGGCUUGAAUUGAAUUAAAUUAAAAUUCAAUGA ...(((.(((..(((.((...........)).))).))).)))((..((((....((((((..((((......))))..))))))..))))..))((((((((.......)))))))).. ( -24.10) >DroSim_CAF1 4003 120 - 1 UAAAAAAAUGUUCCCAGCUUUUUAUUAUCACCGGGCGAUCUUGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUUGGCUUGCAUUGGAUUAAAUUAAAAUUCAAUGA ((((((..((....))..)))))).(((.((...((((........))))...)).)))(((((((((((((........))))))))))))).(((((((((.......))))))))). ( -29.10) >DroPer_CAF1 25257 120 - 1 UACCAAAAUUUUCCCAGCUUUUUAUUAUCGCCGGGUGAUCUUGUCUUGCCAUUGUAAUAGAGUCAGAUGCAAUUUUGUGUUUGUUUUUGGCUUGAAUUGAAUUAAAUUAAAAUUCAAUGA ...(((.(((..(((.((...........)).))).))).)))((..((((....((((((..((((......))))..))))))..))))..))((((((((.......)))))))).. ( -24.10) >DroYak_CAF1 50448 120 - 1 UAAAAAAAUGUUCCCAGUUUUUUAUUAUCACCGGGCGAUCUCGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUUGGCUUGCAUUGGAUUAAAUUAAAAUUCAAUGA (((((((.((....)).)))))))........(((((....))))).............(((((((((((((........))))))))))))).(((((((((.......))))))))). ( -30.50) >consensus UAAAAAAAUGUUCCCAGCUUUUUAUUAUCACCGGGCGAUCUUGUCUUUGCAUUGUAAUAGAGUCAGAUGCAAUUUUUUGUUUGCAUUUGGCUUGCAUUGGAUUAAAUUAAAAUUCAAUGA ............(((.((...........)).)))........................(((((((((((((........))))))))))))).(((((((((.......))))))))). (-22.44 = -23.04 + 0.60)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:10:53 2006