
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,656,678 – 21,656,896 |
| Length | 218 |
| Max. P | 0.938347 |

| Location | 21,656,678 – 21,656,790 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 76.16 |
| Mean single sequence MFE | -30.93 |
| Consensus MFE | -17.08 |
| Energy contribution | -19.70 |
| Covariance contribution | 2.62 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.55 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938347 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21656678 112 + 23771897 AAC-------AGCAAUGAUAUUGUCCUCCUGUAUACAACUGAACAAGUUUUCACAUAUAGCCGAUGGAUCGAGUG-UGACUGCUCUCUAACCAGCAGAGGCAUUCCGUCCGUCGGCCCAA .((-------((....(((...)))...))))....((((.....))))..........((((((((((.(((((-(..(((((........)))))..)))))).)))))))))).... ( -35.70) >DroEre_CAF1 32924 119 + 1 AACAACACC-AGUAAUAAUAUUGUCCUCCUGGAACAAACUGAACAAGUUUUCACAUAUAGCCGAUGGAUCGAGUGCUUACUGCUCUCUAACCAGCAGAGGCAUUCUGUCCGUCGGCCCAA ........(-(((.......((((((....)).))))))))..................((((((((((.((((((((.(((((........))))))))))))).)))))))))).... ( -42.41) >DroYak_CAF1 42058 118 + 1 AACAACACC-AGCAAUGAUAUUGUCAUCCUGUAACUAACUGAACAAGUUUUCACAUAUAGCCGCUAGAUCGAGUG-UGACUGCUCUCUAACCAGCAGAGGCAUUCUGUCCGUCGGCCCAA ........(-((..(((((...))))).))).....((((.....))))..........((((...(((.(((((-(..(((((........)))))..)))))).)))...)))).... ( -26.20) >DroAna_CAF1 33107 110 + 1 GA--ACUUGAUGGAAUAGUAUUGA-------GUACAAACUGAACAAAAUAUCAUUUAUAUCCUAUUCCCCAAGUG-UGUCCAAGCUCUAAGCAGCAGUGGCAUUCUGUCCAUCGGUCCAA ..--(((((..(((((((..((.(-------((....))).))....(((.....)))...))))))).)))))(-.(.((..(((......))).(((((.....).)))).)).)).. ( -19.40) >consensus AAC_ACACC_AGCAAUAAUAUUGUCCUCCUGGAACAAACUGAACAAGUUUUCACAUAUAGCCGAUGGAUCGAGUG_UGACUGCUCUCUAACCAGCAGAGGCAUUCUGUCCGUCGGCCCAA ...........................................................((((((((((.(((((.(..(((((........)))))..)))))).)))))))))).... (-17.08 = -19.70 + 2.62)



| Location | 21,656,711 – 21,656,825 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.04 |
| Mean single sequence MFE | -30.65 |
| Consensus MFE | -21.52 |
| Energy contribution | -24.40 |
| Covariance contribution | 2.88 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.552964 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21656711 114 + 23771897 GAACAAGUUUUCACAUAUAGCCGAUGGAUCGAGUG-UGACUGCUCUCUAACCAGCAGAGGCAUUCCGUCCGUCGGCCCAAACAUGGCAAAAGGCAUGUCC-----AUCCUUCCUAUCGCA ((.................((((((((((.(((((-(..(((((........)))))..)))))).))))))))))....(((((.(....).)))))..-----..........))... ( -38.50) >DroEre_CAF1 32963 115 + 1 GAACAAGUUUUCACAUAUAGCCGAUGGAUCGAGUGCUUACUGCUCUCUAACCAGCAGAGGCAUUCUGUCCGUCGGCCCAAACAUGGCAAAAGGCAUGUCC-----AUCCUUCCUAUCGCA ((.................((((((((((.((((((((.(((((........))))))))))))).))))))))))....(((((.(....).)))))..-----..........))... ( -42.20) >DroYak_CAF1 42097 114 + 1 GAACAAGUUUUCACAUAUAGCCGCUAGAUCGAGUG-UGACUGCUCUCUAACCAGCAGAGGCAUUCUGUCCGUCGGCCCAAACAUGGCAAAAGGCAUGUCC-----AUCCUUCCUAUCGCA ((.................((((...(((.(((((-(..(((((........)))))..)))))).)))...))))....(((((.(....).)))))..-----..........))... ( -28.00) >DroAna_CAF1 33138 119 + 1 GAACAAAAUAUCAUUUAUAUCCUAUUCCCCAAGUG-UGUCCAAGCUCUAAGCAGCAGUGGCAUUCUGUCCAUCGGUCCAAACAAGGCAAAAGGCAUGUCCAUUCCAUCCUUCCUAUCGUA ................(((((((.((((.....((-.(.((..(((......))).(((((.....).)))).)).))).....)).)).))).))))...................... ( -13.90) >consensus GAACAAGUUUUCACAUAUAGCCGAUGGAUCGAGUG_UGACUGCUCUCUAACCAGCAGAGGCAUUCUGUCCGUCGGCCCAAACAUGGCAAAAGGCAUGUCC_____AUCCUUCCUAUCGCA ((.................((((((((((.(((((.(..(((((........)))))..)))))).))))))))))....(((((.(....).))))).................))... (-21.52 = -24.40 + 2.88)

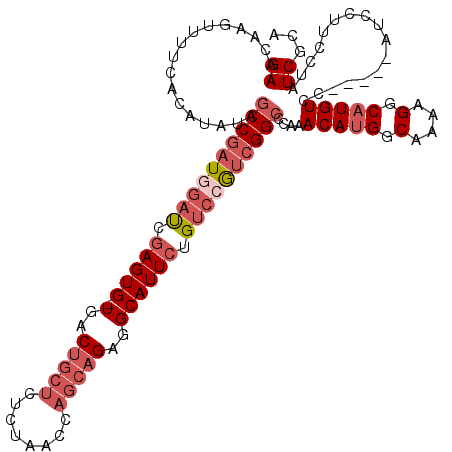

| Location | 21,656,790 – 21,656,896 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.96 |
| Mean single sequence MFE | -31.61 |
| Consensus MFE | -22.17 |
| Energy contribution | -22.57 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.56 |
| SVM RNA-class probability | 0.782167 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21656790 106 + 23771897 ACAUGGCAAAAGGCAUGUCC-----AUCCUUCCUAUCGCAACAAGAGCAAUUUGUUCUAAUGCGCCAUGGCAGCGGCAGCUGCU---GGAAAUGGC-----GAGACU-AAAUCUGUGGAA .((((.(....).))))(((-----((........(((((...(((((.....)))))..)))(((((((((((....))))))---....)))))-----))....-......))))). ( -34.07) >DroVir_CAF1 19607 85 + 1 ACAAGGCAAAAGGCAUGUCC-----AUCC-UCCUAUCGUAACAAGAACAAUUUGUUCUAAUGCGCCAUGGCAGCAGUUGCAGCAGCGGCAG----------------------------- ...(((....(((.((....-----))))-))))...(((((.(((((.....)))))..(((((....)).)))))))).((....))..----------------------------- ( -19.20) >DroEre_CAF1 33043 106 + 1 ACAUGGCAAAAGGCAUGUCC-----AUCCUUCCUAUCGCAACAAGAGCAAUUUGUUCUAAUGCACCAUGGCAGCGGCAGCUGCU---GGAAAUGGC-----GAGACU-AAAUCUGUGGAA .((((.(....).))))(((-----((........((((....(((((.....)))))......(((..(((((....))))))---)).....))-----))....-......))))). ( -30.57) >DroYak_CAF1 42176 106 + 1 ACAUGGCAAAAGGCAUGUCC-----AUCCUUCCUAUCGCAACAAGAGCAAUUUGUUCUAAUGCGCCAUGGCAGCGGCAGCUGCU---GAAAAUGGC-----GAGACU-AAAUGUGUGGAA (((((.(....).)))))..-----.....(((...((((...(((((.....)))))....((((((((((((....))))))---....)))))-----).....-...)))).))). ( -35.00) >DroAna_CAF1 33217 117 + 1 ACAAGGCAAAAGGCAUGUCCAUUCCAUCCUUCCUAUCGUAACAAGAGCAAUUUGUUCUAAUGCGCCAUGGCAGCAGCAGCUGCC---GGAGUUGGGAUGCUGGGCCCAGAAUCUCAAGCU ....(((....(((((.((.(((((...........((((...(((((.....)))))..))))....((((((....))))))---))))).)).)))))..))).............. ( -39.20) >consensus ACAUGGCAAAAGGCAUGUCC_____AUCCUUCCUAUCGCAACAAGAGCAAUUUGUUCUAAUGCGCCAUGGCAGCGGCAGCUGCU___GGAAAUGGC_____GAGACU_AAAUCUGUGGAA .((((((...(((..................)))...(((...(((((.....)))))..)))))))))(((((....)))))..................................... (-22.17 = -22.57 + 0.40)

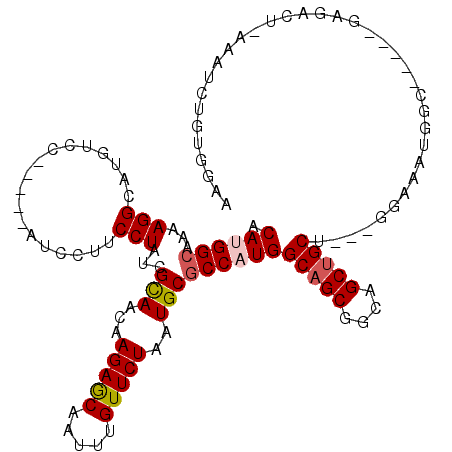
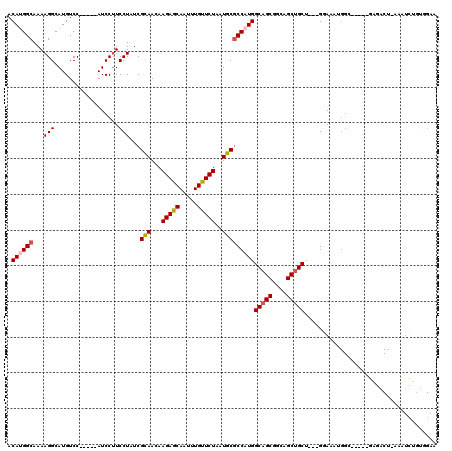
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:10:20 2006