
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,635,016 – 21,635,156 |
| Length | 140 |
| Max. P | 0.825902 |

| Location | 21,635,016 – 21,635,133 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.46 |
| Mean single sequence MFE | -32.58 |
| Consensus MFE | -29.68 |
| Energy contribution | -29.35 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.825902 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21635016 117 - 23771897 CACUCACUGGCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAUAAACUAAACGACAGCUAACGGCGUUGACGCCAAGGAAUGA (((.(((......---..))).)))...((.(((((.(((((..((((((...........))))))..)))))..............((((.((.....))))))..))))).)).... ( -32.20) >DroSec_CAF1 11774 117 - 1 CACUCACUGGCAC---UUAUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAUAAACUAAACGACAGCUAACGGCGUUGACGCCAAGGAAUGA ...((((..((((---....))))))))((.(((((.(((((..((((((...........))))))..)))))..............((((.((.....))))))..))))).)).... ( -32.20) >DroSim_CAF1 11884 117 - 1 CACUCACUGGCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAUAAACUAAACGACAGCUAACGGCGUUGACGCCAAGGAAUGA (((.(((......---..))).)))...((.(((((.(((((..((((((...........))))))..)))))..............((((.((.....))))))..))))).)).... ( -32.20) >DroEre_CAF1 10550 117 - 1 CAUUCACUGCCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAUAAACUAAACGACAGCUAACGGCGUUGACGCCAAGGAAUGA (((((.(((((((---........))))).((((((.(((((..((((((...........))))))..))))).........(.....)...))))))((((....)))).))))))). ( -35.30) >DroYak_CAF1 20883 117 - 1 CAUUCACUGGCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAUAAACUAAACGACAGCUAACGGCGUUGACGCCAAGGAAUGA (((((..((((((---......))...(((((((((.(((((..((((((...........))))))..))))).........(.....)...)))))))))......))))..))))). ( -35.20) >DroAna_CAF1 10053 120 - 1 AACAUAUAUGGAGUGAGUGUCUGUGUGGUCGUUGGCAGUAAGCAGUUCGGCACAUAAUUCACUGAGCAAUUUGCAACAAUAAACUAAACGACAGCUAACGGCGUUGACGCCAAGGAAUGG ..(((.......((.((((((.((.((((.((((...(((((..((((((...........))))))..)))))..))))..)))).)))))).)).))((((....)))).....))). ( -28.40) >consensus CACUCACUGGCAC___UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAUAAACUAAACGACAGCUAACGGCGUUGACGCCAAGGAAUGA ....(((..((((.....))))..))).((.(((((.(((((..((((((...........))))))..)))))..............((((.((.....))))))..))))).)).... (-29.68 = -29.35 + -0.33)

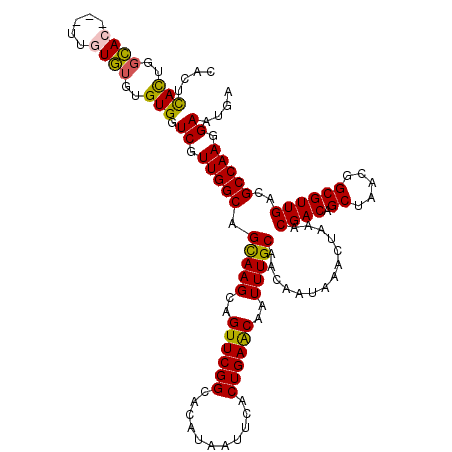

| Location | 21,635,056 – 21,635,156 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.44 |
| Mean single sequence MFE | -27.56 |
| Consensus MFE | -22.18 |
| Energy contribution | -22.07 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.592208 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21635056 100 - 23771897 CCUGCCAAGCCUUUAAA-----------CAU------ACACACUCACUGGCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAU ..(((((((((.....(-----------(((------(((((...........---.))))))))))))..))))))(((((..((((((...........))))))..)))))...... ( -29.10) >DroSec_CAF1 11814 100 - 1 CCUGCCAAGCCUUUAAA-----------CAU------ACACACUCACUGGCAC---UUAUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAU ..(((((((((......-----------...------((((((..........---....)))))))))..))))))(((((..((((((...........))))))..)))))...... ( -25.44) >DroSim_CAF1 11924 100 - 1 CCUGCCAAGCCUUUAAA-----------CAU------ACACACUCACUGGCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAU ..(((((((((.....(-----------(((------(((((...........---.))))))))))))..))))))(((((..((((((...........))))))..)))))...... ( -29.10) >DroEre_CAF1 10590 106 - 1 CCUGCCAAGCCUUUAAA-----------CAUACUCGUACACAUUCACUGCCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAU ..(((((((((.....(-----------(((((..((.((.......))..))---..))))))..)))..))))))(((((..((((((...........))))))..)))))...... ( -26.70) >DroYak_CAF1 20923 100 - 1 CCUGCCAAGCCUUCAAA-----------UAU------ACACAUUCACUGGCAC---UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAU ..(((((((((.....(-----------(((------(((((...........---.))))))))))))..))))))(((((..((((((...........))))))..)))))...... ( -28.10) >DroAna_CAF1 10093 114 - 1 CCUGCCAAGCCCUCGAAAUCAAAAACCACAU------ACAAACAUAUAUGGAGUGAGUGUCUGUGUGGUCGUUGGCAGUAAGCAGUUCGGCACAUAAUUCACUGAGCAAUUUGCAACAAU .(((((((......(....)....(((((((------(((..(((.......)))..)))..)))))))..)))))))...(((((((((...........))))))....)))...... ( -26.90) >consensus CCUGCCAAGCCUUUAAA___________CAU______ACACACUCACUGGCAC___UUGUGUGUGUGGUCGUUGGCAGCAAGCAGUUCGGCACAUAAUUCACUGAACAAUUUGCAACAAU ..(((((((((..........................((((((...............))))))..)))..))))))(((((..((((((...........))))))..)))))...... (-22.18 = -22.07 + -0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:10:02 2006