| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
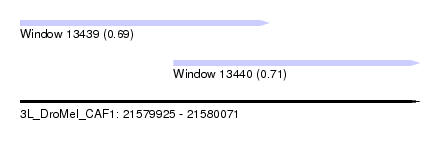
| Location | 21,579,925 – 21,580,071 |
| Length | 146 |
| Max. P | 0.708620 |

| Location | 21,579,925 – 21,580,016 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 82.19 |
| Mean single sequence MFE | -19.88 |
| Consensus MFE | -11.88 |
| Energy contribution | -11.75 |
| Covariance contribution | -0.13 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.691787 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21579925 91 + 23771897 GACAACAAACAAGCUCGUAAGU--UCAAGCACACAA-ACAUAA--------AUAGACACACGUAUGCAAACCUAUUUAUGUGUUUGUAAGAGUUCGAAUGGG ....((((((..(((.(.....--.).)))......-((((((--------((((................))))))))))))))))............... ( -16.29) >DroEre_CAF1 10912 87 + 1 GACAACAAACAAGCACGUAAGU--CUAAGCACACAA-GCAUAA--------AUAAGCACACGUAUGCA----UAUUUAUGUGUUUGUAAGAGUUCGAAUGGG ........((((((((((((((--....((......-))....--------....(((......))).----.))))))))))))))............... ( -18.80) >DroYak_CAF1 8624 87 + 1 UACAACAAACAAGCUCGUAAGU--UCAAGCACACAA-ACAUAA--------AUAAGCACACGUAUGCA----UAUUUAUGUGUUUGUAAGAGAUCGAAUGGG ....((((((..(((.(.....--.).)))......-((((((--------(((.(((......))).----)))))))))))))))............... ( -18.80) >DroAna_CAF1 12781 98 + 1 GACAACAAACAACCUCGUAAAUAGUCAAGCACAUAACACACAAACACUCGGAUAGAUCCGAGUAUGCA----UAUUUAUGUGUUUGUACAAGUUCGAAUGGG ............(((((.((...(((((((((((((.........(((((((....))))))).....----...))))))))))).))...)))))..)). ( -25.63) >consensus GACAACAAACAAGCUCGUAAGU__UCAAGCACACAA_ACAUAA________AUAAACACACGUAUGCA____UAUUUAUGUGUUUGUAAGAGUUCGAAUGGG ........((((((.(((((((......(((.((...........................)).)))......))))))).))))))............... (-11.88 = -11.75 + -0.13)



| Location | 21,579,981 – 21,580,071 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 89.96 |
| Mean single sequence MFE | -18.77 |
| Consensus MFE | -16.62 |
| Energy contribution | -15.88 |
| Covariance contribution | -0.75 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.708620 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21579981 90 + 23771897 AAACCUAUUUAUGUGUUUGUAAGAGUUCGAAUGGGUUUAUGUAUAUUCACCCAUGGCUGAAGUAUUUUUAUUGACUUCAGCUUUGGCGUU ((((((((((..(..(((....)))..)))))))))))..........(((((.(((((((((..........))))))))).))).)). ( -23.90) >DroEre_CAF1 10968 86 + 1 A----UAUUUAUGUGUUUGUAAGAGUUCGAAUGGGUUUAUGUAUAUUCACCCAUGGUUGAAGUAUUUUUAUUGACUUUAGCUUUGGCGUU .----.......(((..((((.(((..(....)..)))...))))..)))(((.(((((((((..........))))))))).))).... ( -17.50) >DroYak_CAF1 8680 85 + 1 A----UAUUUAUGUGUUUGUAAGAGAUCGAAUGGGUUUAUGUAUAUUCACCCAUGGUUGAGGUGUUUUUAUUGACUUUAGCUUUGGCGU- .----.......(((..((((..(((((.....)))))...))))..)))(((.(((((((((..........))))))))).)))...- ( -19.10) >DroAna_CAF1 12848 86 + 1 A----UAUUUAUGUGUUUGUACAAGUUCGAAUGGGCUUAUGUAUAGUCACCCAUGGGUAAACUAUUUUUAUUGACUUUGGCUUUGGCGUU .----((((((((.((((((((((((((....)))))..)))))))..)).))))))))....................((....))... ( -14.60) >consensus A____UAUUUAUGUGUUUGUAAGAGUUCGAAUGGGUUUAUGUAUAUUCACCCAUGGUUGAAGUAUUUUUAUUGACUUUAGCUUUGGCGUU ............(((..((((.(((..(....)..)))...))))..)))(((.(((((((((..........))))))))).))).... (-16.62 = -15.88 + -0.75)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:09:37 2006