
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 2,488,896 – 2,488,987 |
| Length | 91 |
| Max. P | 0.999912 |

| Location | 2,488,896 – 2,488,987 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 93.46 |
| Mean single sequence MFE | -21.19 |
| Consensus MFE | -17.37 |
| Energy contribution | -16.89 |
| Covariance contribution | -0.48 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.82 |
| SVM decision value | 2.87 |
| SVM RNA-class probability | 0.997506 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 2488896 91 + 23771897 AUAAUAAUUGGCUAGUCGCAUUAAUUGCUAAGUGUAGUGUUUCUUUUAUAAA--CCUUCACAAUACUUUUUGCGCUGACCAAUUCCGUUUUUA .....((((((.(((.((((.........((((((.(((.............--....))).))))))..))))))).))))))......... ( -17.23) >DroSec_CAF1 112337 91 + 1 AUAAUAAUUGGCUAGUCGCAUUAAUUGCUAAGUGUAGUGUUUCUUCUAUAAA--CCCUCACAAUACUUUUUGCGCUAACCAAUUCUGUUUUUA .....((((((.(((.((((.........((((((.(((.............--....))).))))))..))))))).))))))......... ( -17.53) >DroSim_CAF1 107277 91 + 1 AUAAUAAUUGGCUAGUCGCAUUAAUUGCUAAGUGUAGUGUUUCUUCUAUGAA--CCCUCACAAUACUUUUUGCGCUGACCAAUUCUGUUUUUA .....((((((.(((.((((.........((((((.(((.(((......)))--....))).))))))..))))))).))))))......... ( -19.40) >DroEre_CAF1 112562 91 + 1 AUAAUAAUUGGCUAGUCGCAUUAAUUGCUAAGUGUAGUGUUUCUUCUAUAAA--CCUUCGCAAUACUUUUUGCGCUGGCCAAUUCUGUUUUUA .....((((((((((.((((...(((((.((((((((........)))))..--.))).)))))......))))))))))))))......... ( -27.00) >DroYak_CAF1 110615 93 + 1 AUAAUAAUUGGCUAGUCGCAUUAAUUGCCAAGUGUAGUGUUUCUUCUAUACUUUUCUUCACAAUACUUUUUGCGCUGGCCGAUUCUGUUUUUA .....((((((((((.((((...((((..(((...(((((.......)))))...)))..))))......))))))))))))))......... ( -24.80) >consensus AUAAUAAUUGGCUAGUCGCAUUAAUUGCUAAGUGUAGUGUUUCUUCUAUAAA__CCUUCACAAUACUUUUUGCGCUGACCAAUUCUGUUUUUA .....((((((.(((.((((.........((((((.(((...................))).))))))..))))))).))))))......... (-17.37 = -16.89 + -0.48)

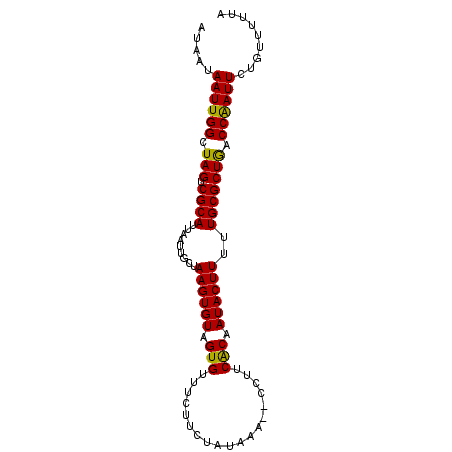
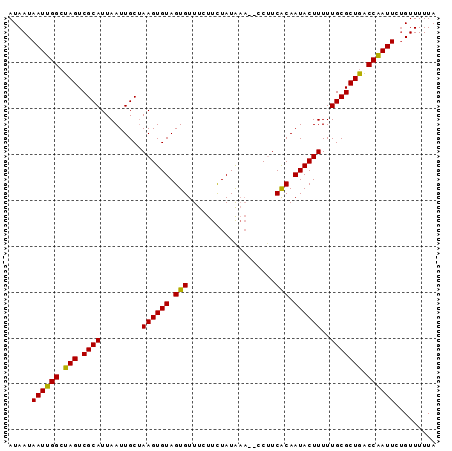
| Location | 2,488,896 – 2,488,987 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 93.46 |
| Mean single sequence MFE | -20.18 |
| Consensus MFE | -18.38 |
| Energy contribution | -17.94 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.91 |
| SVM decision value | 4.51 |
| SVM RNA-class probability | 0.999912 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 2488896 91 - 23771897 UAAAAACGGAAUUGGUCAGCGCAAAAAGUAUUGUGAAGG--UUUAUAAAAGAAACACUACACUUAGCAAUUAAUGCGACUAGCCAAUUAUUAU .........((((((..((((((....((...(((.(((--(((.......)))).)).)))...))......)))).))..))))))..... ( -19.30) >DroSec_CAF1 112337 91 - 1 UAAAAACAGAAUUGGUUAGCGCAAAAAGUAUUGUGAGGG--UUUAUAGAAGAAACACUACACUUAGCAAUUAAUGCGACUAGCCAAUUAUUAU .........((((((((((((((....((...(((.(((--(((.......)))).)).)))...))......)))).))))))))))..... ( -22.70) >DroSim_CAF1 107277 91 - 1 UAAAAACAGAAUUGGUCAGCGCAAAAAGUAUUGUGAGGG--UUCAUAGAAGAAACACUACACUUAGCAAUUAAUGCGACUAGCCAAUUAUUAU .........((((((..((((((....((...(((((..--(((......)))...)).)))...))......)))).))..))))))..... ( -19.40) >DroEre_CAF1 112562 91 - 1 UAAAAACAGAAUUGGCCAGCGCAAAAAGUAUUGCGAAGG--UUUAUAGAAGAAACACUACACUUAGCAAUUAAUGCGACUAGCCAAUUAUUAU .........(((((((.((((((......(((((..(((--(...(((........))).)))).)))))...)))).)).)))))))..... ( -21.60) >DroYak_CAF1 110615 93 - 1 UAAAAACAGAAUCGGCCAGCGCAAAAAGUAUUGUGAAGAAAAGUAUAGAAGAAACACUACACUUGGCAAUUAAUGCGACUAGCCAAUUAUUAU .............(((.((((((......(((((.(((...(((...........)))...))).)))))...)))).)).)))......... ( -17.90) >consensus UAAAAACAGAAUUGGUCAGCGCAAAAAGUAUUGUGAAGG__UUUAUAGAAGAAACACUACACUUAGCAAUUAAUGCGACUAGCCAAUUAUUAU .........(((((((.((((((......(((((.(((.......................))).)))))...)))).)).)))))))..... (-18.38 = -17.94 + -0.44)

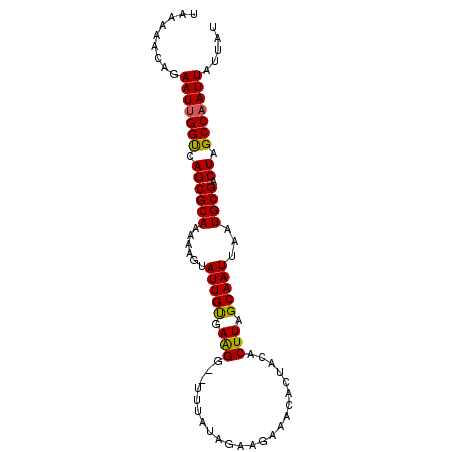
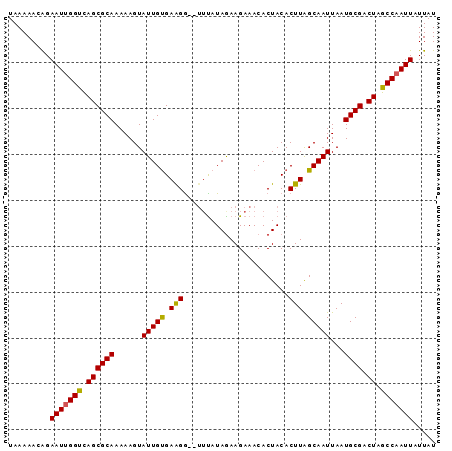
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:51:54 2006