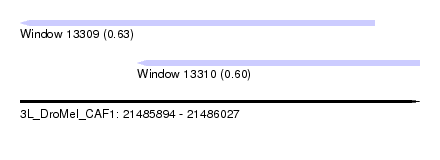
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,485,894 – 21,486,027 |
| Length | 133 |
| Max. P | 0.632773 |

| Location | 21,485,894 – 21,486,012 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 86.27 |
| Mean single sequence MFE | -22.80 |
| Consensus MFE | -17.81 |
| Energy contribution | -18.88 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.632773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
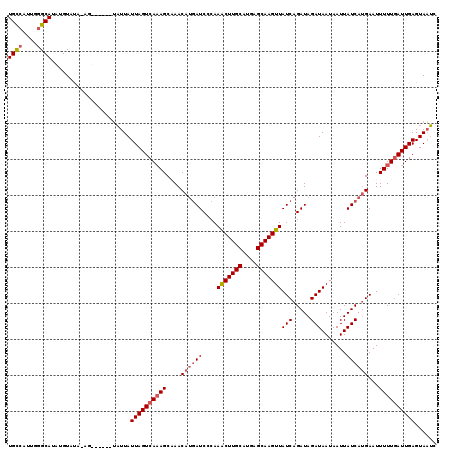
>3L_DroMel_CAF1 21485894 118 - 23771897 UGCCAUUGGGCAUAUGUAUA-AGUAUUAUUAUUAUUAGUCAAAGCAAACAUGAUCACAAGCUUGCAUAAGCAAGUUAUCAGAUAGAUAAUAAUUAUCAUGAAUUUUUGAUUGAGUAAUC ((((....))))........-......(((((..((((((((((....((((((.....((((((....))))))((((.....))))......))))))...))))))))))))))). ( -25.20) >DroSec_CAF1 7696 112 - 1 UGUCAUUGGGCAUAUGUAUU-AG------UAUUAUUAGUCAAAGCUAACAUGAUCCCAAACUUGCAUGAGCAAGUUAUCAGAUAGAUAAUAAUUAUCAUGAAUUUUUAAUUGAGUAAUC ((((..((((..(((((..(-((------(.............)))))))))..))))(((((((....)))))))....))))((((.....))))...................... ( -20.72) >DroSim_CAF1 9085 113 - 1 UGUCAUUGGGCAUUUGUAUUAAG------UAUUAUUAGUCAAAGCUAACAUGAUCCCAAACUUGCAUGAGCAAGUUAUCAGAUAGAUAAUAAUUAUCAUGAAUUUUUGAUUGAGUAAUC ((((....))))...........------.....((((((((((....((((((....(((((((....)))))))(((.....))).......))))))...))))))))))...... ( -22.20) >DroYak_CAF1 9564 114 - 1 UGCCAAUAAGCAUAUGUACA-UAUAU---GUUUAUUAGUCACAGCAAACAGUAUCCCAAACUUGCAUGAGCAAGUUAUCAGAUAGAUAAUAAUUAUCAAGAAUUUUUGAUUGAGUACA- .((.((((((((((((....-)))))---))))))).))..(((((((...((((...(((((((....)))))))....))))((((.....)))).......)))).)))......- ( -23.10) >consensus UGCCAUUGGGCAUAUGUAUA_AG______UAUUAUUAGUCAAAGCAAACAUGAUCCCAAACUUGCAUGAGCAAGUUAUCAGAUAGAUAAUAAUUAUCAUGAAUUUUUGAUUGAGUAAUC ((((....))))......................((((((((((....((((((....(((((((....)))))))(((.....))).......))))))...))))))))))...... (-17.81 = -18.88 + 1.06)
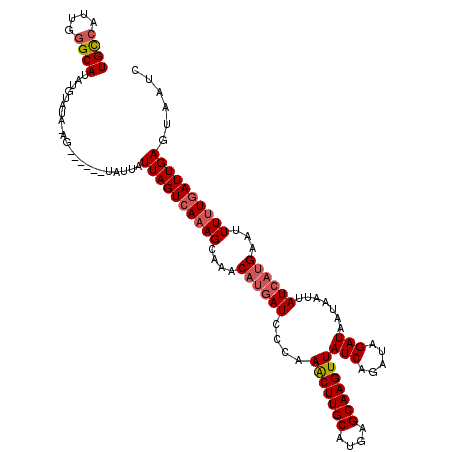


| Location | 21,485,933 – 21,486,027 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 79.48 |
| Mean single sequence MFE | -19.54 |
| Consensus MFE | -11.26 |
| Energy contribution | -11.26 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.600606 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
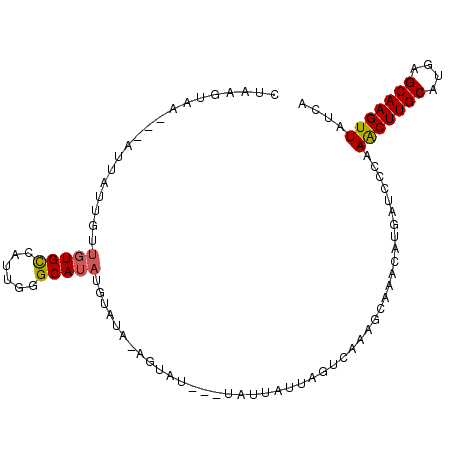
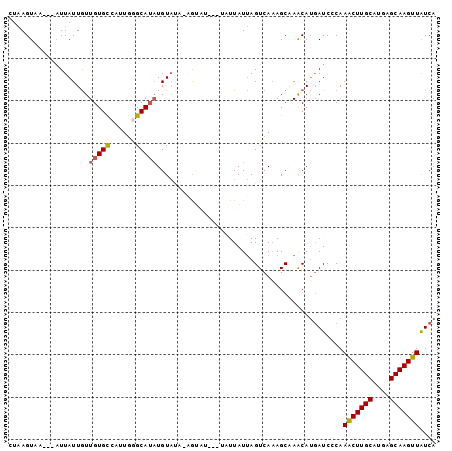
>3L_DroMel_CAF1 21485933 94 - 23771897 UUAAGU------UUAUUGUUGUGCCAUUGGGCAUAUGUAUA-AGUAUUAUUAUUAUUAGUCAAAGCAAACAUGAUCACAAGCUUGCAUAAGCAAGUUAUCA .....(------((((...((((((....))))))...)))-))...........................((((....(((((((....))))))))))) ( -17.50) >DroSec_CAF1 7735 91 - 1 CUAAGUCA---AUUAUUGUUGUGUCAUUGGGCAUAUGUAUU-AG------UAUUAUUAGUCAAAGCUAACAUGAUCCCAAACUUGCAUGAGCAAGUUAUCA ......((---((....))))......((((..(((((..(-((------(.............)))))))))..))))(((((((....))))))).... ( -17.22) >DroSim_CAF1 9124 92 - 1 CUAAGUCA---AUUAUUGUUGUGUCAUUGGGCAUUUGUAUUAAG------UAUUAUUAGUCAAAGCUAACAUGAUCCCAAACUUGCAUGAGCAAGUUAUCA ......((---((....)))).(((((..(((.((((.(((((.------.....))))))))))))...)))))....(((((((....))))))).... ( -18.70) >DroEre_CAF1 7819 90 - 1 ---AGUAAAUAAUUAUUAUUGUGCCAAUGAGCAAAUGU-----AUAU---GUUUAUAAGUCAAAGCAAGCAUGAUCCCAAACUUGCAUGAGCAAGUUGUCA ---..............((..(((..(((((((.....-----...)---))))))..((....))..)))..))....(((((((....))))))).... ( -16.10) >DroYak_CAF1 9602 94 - 1 UUAAGUAA---AUUAUUGUUGUGCCAAUAAGCAUAUGUACA-UAUAU---GUUUAUUAGUCACAGCAAACAGUAUCCCAAACUUGCAUGAGCAAGUUAUCA ........---....(((((((((.((((((((((((....-)))))---))))))).).))))))))...........(((((((....))))))).... ( -28.20) >consensus CUAAGUAA___AUUAUUGUUGUGCCAUUGGGCAUAUGUAUA_AGUAU___UAUUAUUAGUCAAAGCAAACAUGAUCCCAAACUUGCAUGAGCAAGUUAUCA ...................(((((......)))))............................................(((((((....))))))).... (-11.26 = -11.26 + -0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:07:29 2006