
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,414,090 – 21,414,196 |
| Length | 106 |
| Max. P | 0.801814 |

| Location | 21,414,090 – 21,414,196 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 83.08 |
| Mean single sequence MFE | -41.00 |
| Consensus MFE | -23.73 |
| Energy contribution | -26.23 |
| Covariance contribution | 2.50 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.801814 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
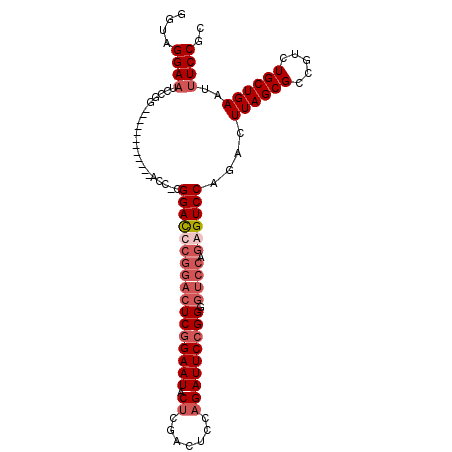
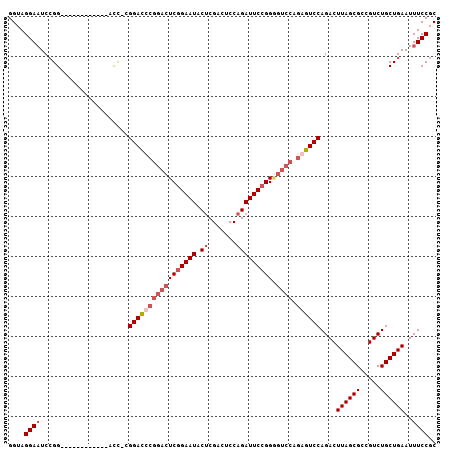
>3L_DroMel_CAF1 21414090 106 + 23771897 GCGGAAAUUCAGCAGACGGCGCUAAGUCUGGACUCUGGACUCCGGAAUCUGGAGUCGAGUAUUCGGAGUCCGGGUCCG-GGUCCGGGUCCGGAUCCGGAUUCCUACC ..(((.((((..(((((........))))).......((((((((...)))))))))))).)))((((((((((((((-((......)))))))))))))))).... ( -55.10) >DroSec_CAF1 5488 94 + 1 GCGGAAAUUCAGCAGACGGCGCUAAGUCUGGACUCUGGACCCCGGAAUCUGGAGUCGAGUAUUCCGAGUCCGAGUCCG-GGU------------CCGGAUUCCUACC ..((((......(((((........)))))...(((((((((.(((.((.(((.(((.......))).))))).))))-)))------------))))))))).... ( -41.70) >DroSim_CAF1 5692 94 + 1 GCGGAAAUUCAGCAGACGGCGCUAAGUCUGGACUCUGGACCCCGGAAUCUGGAGUCGAGUAUUCCGAGUCCGAGUCCG-GGU------------CCGGAUUCCUACC ..((((......(((((........)))))...(((((((((.(((.((.(((.(((.......))).))))).))))-)))------------))))))))).... ( -41.70) >DroYak_CAF1 5548 88 + 1 GCGGAAAUUCAGCAGACGGCGCUAAGUCUGGACU-------CCGGAAUCUGGAGCCGCGUAUUCCGAAGCUGGAUCCUGAAU------------CCUGAAUCCUACC .(((((.....((((((........))))((.((-------((((...))))))))))...))))).....((((.(.....------------...).)))).... ( -25.50) >consensus GCGGAAAUUCAGCAGACGGCGCUAAGUCUGGACUCUGGACCCCGGAAUCUGGAGUCGAGUAUUCCGAGUCCGAGUCCG_GGU____________CCGGAUUCCUACC .(((((((((..(((((........)))))(((((((((........))))))))))))).))))).....(((((((.................)))))))..... (-23.73 = -26.23 + 2.50)



| Location | 21,414,090 – 21,414,196 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 83.08 |
| Mean single sequence MFE | -36.38 |
| Consensus MFE | -23.81 |
| Energy contribution | -26.12 |
| Covariance contribution | 2.31 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.583999 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21414090 106 - 23771897 GGUAGGAAUCCGGAUCCGGACCCGGACC-CGGACCCGGACUCCGAAUACUCGACUCCAGAUUCCGGAGUCCAGAGUCCAGACUUAGCGCCGUCUGCUGAAUUUCCGC ....(((((((((.((((....)))).)-))))...((((((.((....))((((((.......))))))..))))))....((((((.....))))))..)))).. ( -43.20) >DroSec_CAF1 5488 94 - 1 GGUAGGAAUCCGG------------ACC-CGGACUCGGACUCGGAAUACUCGACUCCAGAUUCCGGGGUCCAGAGUCCAGACUUAGCGCCGUCUGCUGAAUUUCCGC ((..(((.((.((------------(((-((((.(((((.((((.....)))).))).)).))).)))))).)).)))....((((((.....))))))....)).. ( -36.50) >DroSim_CAF1 5692 94 - 1 GGUAGGAAUCCGG------------ACC-CGGACUCGGACUCGGAAUACUCGACUCCAGAUUCCGGGGUCCAGAGUCCAGACUUAGCGCCGUCUGCUGAAUUUCCGC ((..(((.((.((------------(((-((((.(((((.((((.....)))).))).)).))).)))))).)).)))....((((((.....))))))....)).. ( -36.50) >DroYak_CAF1 5548 88 - 1 GGUAGGAUUCAGG------------AUUCAGGAUCCAGCUUCGGAAUACGCGGCUCCAGAUUCCGG-------AGUCCAGACUUAGCGCCGUCUGCUGAAUUUCCGC ((...((((((((------------((((.(((((.((((.((.....)).))))...)).))).)-------))))(((((........)))))))))))).)).. ( -29.30) >consensus GGUAGGAAUCCGG____________ACC_CGGACCCGGACUCGGAAUACUCGACUCCAGAUUCCGGGGUCCAGAGUCCAGACUUAGCGCCGUCUGCUGAAUUUCCGC ....((((......................(((((((((((((((((.((.......))))))))).)))).))))))....((((((.....))))))..)))).. (-23.81 = -26.12 + 2.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:07:01 2006