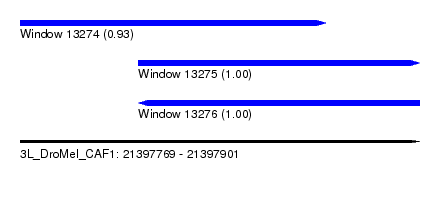

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,397,769 – 21,397,901 |
| Length | 132 |
| Max. P | 0.999282 |

| Location | 21,397,769 – 21,397,870 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 79.62 |
| Mean single sequence MFE | -27.80 |
| Consensus MFE | -19.06 |
| Energy contribution | -18.70 |
| Covariance contribution | -0.36 |
| Combinations/Pair | 1.35 |
| Mean z-score | -2.20 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.927837 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21397769 101 + 23771897 CGCUUUUAAUAAUCUCACUGAUGAUGGCUUCUUGUGGA-UGCAGAUAAGUAUCUCGCCCAAGUGGCUGCUAUUCGUC-AGAUACUUAUCAAUUAAUGUUGAAA .................(((((((.((((.((((.((.-((.((((....)))))))))))).)))).....)))))-)).......(((((....))))).. ( -24.90) >DroSec_CAF1 30454 91 + 1 CGUUUUUAAUAAUCUCG----------CUUCUUGUGGA-UGCAGAUAAGUAUCUCGCUCGAGUGGCUGCCAUUCGUC-GGAUACUUAUCAAUUAAUGCUGAAA .((..(((((.(((.((----------(.....)))))-)...((((((((((.((..(((((((...))))))).)-))))))))))).))))).))..... ( -29.00) >DroSim_CAF1 30558 101 + 1 CGUUUUUAAUAAUCUCGCUGAUGAUGGCUUCUUGUGGA-UGCAGAUAAGUAUCUCGCUCGAGUGGCUGCCAUUCGUC-GGAUACUUAUCAAUUAAUGCUGAAA .((..(((((.(((.(((.((........))..)))))-)...((((((((((.((..(((((((...))))))).)-))))))))))).))))).))..... ( -29.30) >DroEre_CAF1 31554 99 + 1 CGUUUUUAAUAAUCUCGCUGAUGAUGGCUUCUUGUGGG-UACAGAUAAGUAU--CGCUCGAGUGGCUGCCAUUAGGC-GGAUACUUAUCAAUUAAUGCUGGAA ...........(((.((((((((..((((.((((.(((-(((......))))--).).)))).))))..))))).))-))))..................... ( -26.40) >DroYak_CAF1 31569 100 + 1 CGUUUUUCAUAAUCUCGCUGAUGAUGGCUUCUUGUGGA-UACAGAUAAGCAUCUCGCUCGAGUGGCUGCCAUUCGUG-GGAUGCUUAUCAAUUAAUGCUA-AA .((..(((((((....(((......)))...)))))))-.)).((((((((((((((..((((((...)))))))))-)))))))))))...........-.. ( -34.60) >DroPer_CAF1 17258 86 + 1 GGUUUUUAAUAAUCUCGCUGA---UAGCUACUUGUGGCCCGUUGAUAAGUAUCUCACUCAAGUGGCCUGCAUGUGCCGGGAUACUUAUU-------------- ((.........(((.....))---).((((....))))))...((((((((((((((....))((((.....).)))))))))))))))-------------- ( -22.60) >consensus CGUUUUUAAUAAUCUCGCUGAUGAUGGCUUCUUGUGGA_UGCAGAUAAGUAUCUCGCUCGAGUGGCUGCCAUUCGUC_GGAUACUUAUCAAUUAAUGCUGAAA .((.....((((....(((......)))...)))).....)).(((((((((((....((((((.....))))))...))))))))))).............. (-19.06 = -18.70 + -0.36)

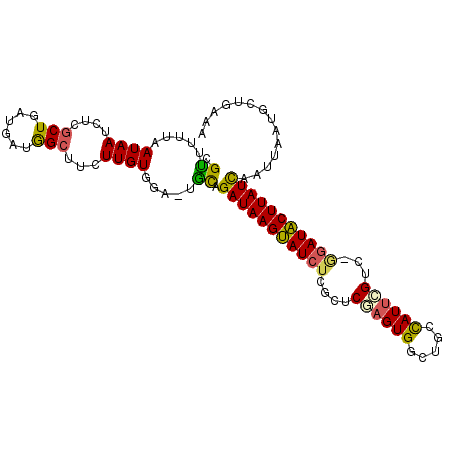
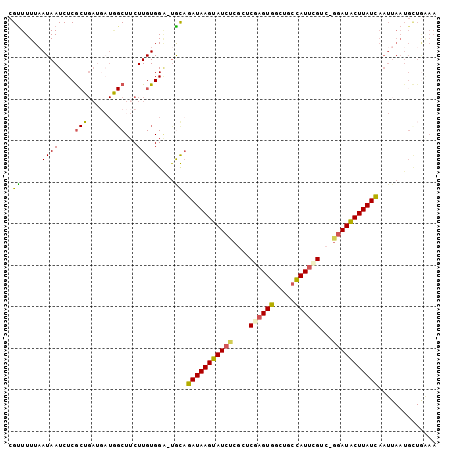
| Location | 21,397,808 – 21,397,901 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 82.90 |
| Mean single sequence MFE | -29.18 |
| Consensus MFE | -22.40 |
| Energy contribution | -22.28 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.22 |
| Mean z-score | -4.45 |
| Structure conservation index | 0.77 |
| SVM decision value | 3.48 |
| SVM RNA-class probability | 0.999282 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21397808 93 + 23771897 GCAGAUAAGUAUCUCGCCCAAGUGGCUGCUAUUCGUCAGAUACUUAUCAAUUAAUGUUGAAAUUAUAGCACUUUAAGUAGCAACAUAAUUCAA ...(((((((((((.((...(((....)))....)).))))))))))).....((((((...(((........)))....))))))....... ( -21.30) >DroSec_CAF1 30483 93 + 1 GCAGAUAAGUAUCUCGCUCGAGUGGCUGCCAUUCGUCGGAUACUUAUCAAUUAAUGCUGAAAUUAUAGCACUAUAAGUGGCAACCUAAUUUAG ...((((((((((.((..(((((((...))))))).))))))))))))......(((((...(((((....))))).)))))........... ( -30.00) >DroSim_CAF1 30597 93 + 1 GCAGAUAAGUAUCUCGCUCGAGUGGCUGCCAUUCGUCGGAUACUUAUCAAUUAAUGCUGAAAUUAUAGCACUAUAAGUGGCAACACAAUUUAG ...((((((((((.((..(((((((...))))))).))))))))))))......(((((......)))))......(((....)))....... ( -32.60) >DroEre_CAF1 31593 91 + 1 ACAGAUAAGUAU--CGCUCGAGUGGCUGCCAUUAGGCGGAUACUUAUCAAUUAAUGCUGGAAUUAUAGCAUUAUAAGUAGCAACAUAAUUUCA ...(((((((((--((((.....)))((((....))))))))))))))...((((((((......)))))))).................... ( -27.30) >DroYak_CAF1 31608 77 + 1 ACAGAUAAGCAUCUCGCUCGAGUGGCUGCCAUUCGUGGGAUGCUUAUCAAUUAAUGCUA-AAUUAUAGCAUUAUAAGA--------------- ...((((((((((((((..((((((...))))))))))))))))))))...((((((((-.....)))))))).....--------------- ( -34.70) >consensus GCAGAUAAGUAUCUCGCUCGAGUGGCUGCCAUUCGUCGGAUACUUAUCAAUUAAUGCUGAAAUUAUAGCACUAUAAGUAGCAACAUAAUUUAA ...(((((((((((....(((((((...)))))))..)))))))))))......(((((......)))))....................... (-22.40 = -22.28 + -0.12)
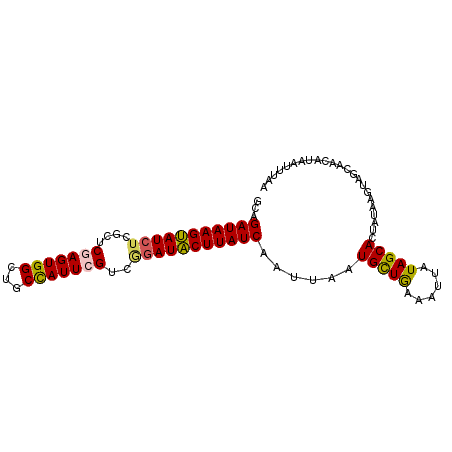


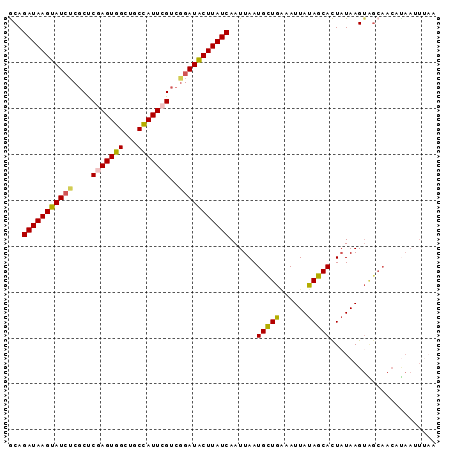
| Location | 21,397,808 – 21,397,901 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 82.90 |
| Mean single sequence MFE | -25.36 |
| Consensus MFE | -18.26 |
| Energy contribution | -18.62 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.16 |
| Mean z-score | -3.33 |
| Structure conservation index | 0.72 |
| SVM decision value | 2.72 |
| SVM RNA-class probability | 0.996591 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21397808 93 - 23771897 UUGAAUUAUGUUGCUACUUAAAGUGCUAUAAUUUCAACAUUAAUUGAUAAGUAUCUGACGAAUAGCAGCCACUUGGGCGAGAUACUUAUCUGC ((((((((((..(((......)))..))))).)))))........((((((((((((.(.....)).(((.....))).)))))))))))... ( -22.10) >DroSec_CAF1 30483 93 - 1 CUAAAUUAGGUUGCCACUUAUAGUGCUAUAAUUUCAGCAUUAAUUGAUAAGUAUCCGACGAAUGGCAGCCACUCGAGCGAGAUACUUAUCUGC ......(((((....)))))(((((((........)))))))...((((((((((((.(((.(((...))).)))..)).))))))))))... ( -26.30) >DroSim_CAF1 30597 93 - 1 CUAAAUUGUGUUGCCACUUAUAGUGCUAUAAUUUCAGCAUUAAUUGAUAAGUAUCCGACGAAUGGCAGCCACUCGAGCGAGAUACUUAUCUGC .......(((....)))...(((((((........)))))))...((((((((((((.(((.(((...))).)))..)).))))))))))... ( -26.20) >DroEre_CAF1 31593 91 - 1 UGAAAUUAUGUUGCUACUUAUAAUGCUAUAAUUCCAGCAUUAAUUGAUAAGUAUCCGCCUAAUGGCAGCCACUCGAGCG--AUACUUAUCUGU ....................(((((((........)))))))...((((((((((.(((....))).(((....).)))--)))))))))... ( -24.50) >DroYak_CAF1 31608 77 - 1 ---------------UCUUAUAAUGCUAUAAUU-UAGCAUUAAUUGAUAAGCAUCCCACGAAUGGCAGCCACUCGAGCGAGAUGCUUAUCUGU ---------------.....((((((((.....-))))))))...((((((((((((.(((.(((...))).))).).).))))))))))... ( -27.70) >consensus CUAAAUUAUGUUGCCACUUAUAGUGCUAUAAUUUCAGCAUUAAUUGAUAAGUAUCCGACGAAUGGCAGCCACUCGAGCGAGAUACUUAUCUGC ....................(((((((........)))))))...((((((((((((.(((.(((...))).)))..)).))))))))))... (-18.26 = -18.62 + 0.36)

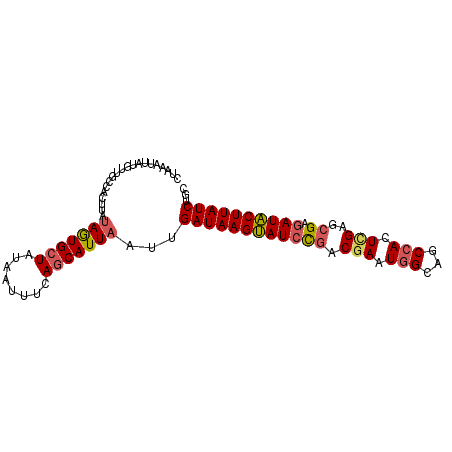

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:06:52 2006