
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,233,412 – 21,233,513 |
| Length | 101 |
| Max. P | 0.632858 |

| Location | 21,233,412 – 21,233,513 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 84.29 |
| Mean single sequence MFE | -23.72 |
| Consensus MFE | -15.54 |
| Energy contribution | -16.64 |
| Covariance contribution | 1.10 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.632858 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
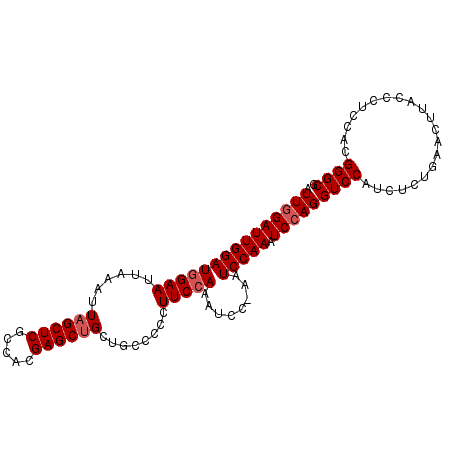
>3L_DroMel_CAF1 21233412 101 + 23771897 AUUGGAUUGGAUGGAAUUAAAUUAGCUCGCCACGAGCUGCUGCCCCCUUCCAAAUCCAAAUCCAAAUCCAGGUCCAUCUCUAAACUUACCAUCCACGGGCG .(((((((((((((((......(((((((...)))))))........)))))..))).)))))))......((((.....................)))). ( -25.84) >DroSec_CAF1 628 101 + 1 AUUGGAUUGGAUGGAAUUAAAUUAGCUCGCCACGAGCAGCGGCCCACUUCCAAAUCCUAAUCCAAAUCCAGGUCCAUCUCUGAACUUACCCUCCACGGGCG .(((((((((((((((........(((((...)))))...(.....))))))..))).)))))))...((((......))))......(((.....))).. ( -25.90) >DroSim_CAF1 638 101 + 1 AUUGGAUUGGAUGGAAUUAAAUUAGCUCUCCACGAGCUGCUGCCCACUUCCAAAUCCUAAUCCAAAUCCAGGUCCAUCUCUGAACUUACCCUCCACGGGCG .(((((((((((((((......((((((.....))))))........)))))..))).)))))))...((((......))))......(((.....))).. ( -27.04) >DroEre_CAF1 586 84 + 1 AUUGGA-----UUGAAUUAAAUUAGCUCGCCACGAGCUGCUGUCCCCUUCCAA------------AUCCAGGUCCAUUUCUGAACUUACCCUCCAUGGGCG .(((((-----(((((......(((((((...)))))))........)))..)------------))))))((((((....((........)).)))))). ( -18.94) >DroYak_CAF1 585 84 + 1 AUUAGA-----UGGAAUUAAAUUAGCUCGCCACGAGCUGCUGCCCCCUUCCAA------------AUCCAGGUCCAUCUCUGAACUUACCCUCUAUGGGCG ...(((-----((((.......(((((((...)))))))......(((.....------------....)))))))))).........(((.....))).. ( -20.90) >consensus AUUGGAUUGGAUGGAAUUAAAUUAGCUCGCCACGAGCUGCUGCCCCCUUCCAAAUCC_AAUCCAAAUCCAGGUCCAUCUCUGAACUUACCCUCCACGGGCG .(((((((((((((((......((((((.....))))))........)))))........))))).)))))((((.....................)))). (-15.54 = -16.64 + 1.10)


| Location | 21,233,412 – 21,233,513 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 84.29 |
| Mean single sequence MFE | -29.96 |
| Consensus MFE | -21.16 |
| Energy contribution | -21.78 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.71 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.522483 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
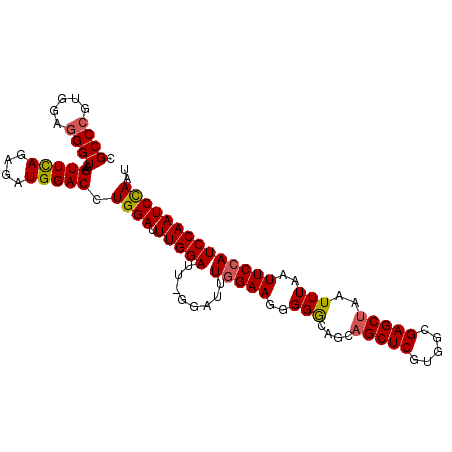
>3L_DroMel_CAF1 21233412 101 - 23771897 CGCCCGUGGAUGGUAAGUUUAGAGAUGGACCUGGAUUUGGAUUUGGAUUUGGAAGGGGGCAGCAGCUCGUGGCGAGCUAAUUUAAUUCCAUCCAAUCCAAU ...(((.....(((..(((....)))..)))......)))..(((((((.(((.((((.....((((((...)))))).......)))).)))))))))). ( -28.10) >DroSec_CAF1 628 101 - 1 CGCCCGUGGAGGGUAAGUUCAGAGAUGGACCUGGAUUUGGAUUAGGAUUUGGAAGUGGGCCGCUGCUCGUGGCGAGCUAAUUUAAUUCCAUCCAAUCCAAU .((((.....))))..(((((....))))).((((((.((((..(((.((((((((..(((((.....)))))..)))..))))).))))))))))))).. ( -36.50) >DroSim_CAF1 638 101 - 1 CGCCCGUGGAGGGUAAGUUCAGAGAUGGACCUGGAUUUGGAUUAGGAUUUGGAAGUGGGCAGCAGCUCGUGGAGAGCUAAUUUAAUUCCAUCCAAUCCAAU .((((.....))))((((((((........))))))))(((((.(((..(((((((.....))(((((.....))))).......)))))))))))))... ( -33.40) >DroEre_CAF1 586 84 - 1 CGCCCAUGGAGGGUAAGUUCAGAAAUGGACCUGGAU------------UUGGAAGGGGACAGCAGCUCGUGGCGAGCUAAUUUAAUUCAA-----UCCAAU .((((.....))))..(((((....))))).(((((------------(.(((.(....)...((((((...)))))).......)))))-----)))).. ( -27.30) >DroYak_CAF1 585 84 - 1 CGCCCAUAGAGGGUAAGUUCAGAGAUGGACCUGGAU------------UUGGAAGGGGGCAGCAGCUCGUGGCGAGCUAAUUUAAUUCCA-----UCUAAU .((((.....))))........((((((((((....------------.....))).......((((((...))))))........))))-----)))... ( -24.50) >consensus CGCCCGUGGAGGGUAAGUUCAGAGAUGGACCUGGAUUUGGAUU_GGAUUUGGAAGGGGGCAGCAGCUCGUGGCGAGCUAAUUUAAUUCCAUCCAAUCCAAU .((((.....))))..(((((....))))).((((.(((((........(((((..(((....(((((.....)))))..)))..)))))))))))))).. (-21.16 = -21.78 + 0.62)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:05:05 2006