
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,162,538 – 21,162,669 |
| Length | 131 |
| Max. P | 0.998511 |

| Location | 21,162,538 – 21,162,653 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.85 |
| Mean single sequence MFE | -29.08 |
| Consensus MFE | -12.05 |
| Energy contribution | -12.55 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.41 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.932654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
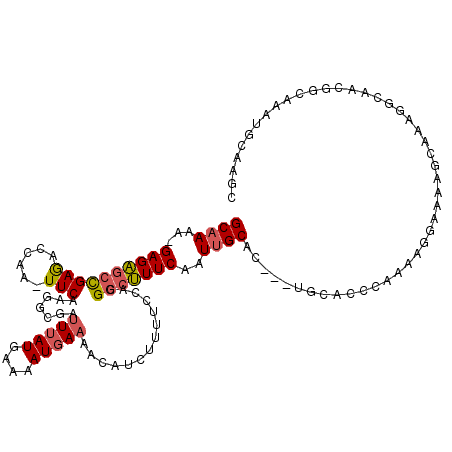
>3L_DroMel_CAF1 21162538 115 - 23771897 GCAAAA-GAGAGCCGAGACCAA-UUCAAGGCGAUUUAUGAAAAUUAAAACAUCUUUUCCAUGGCUUUCAAUUGCAC---UGCACCCAAAAGGAAAAGCAAAUGCAGCGGCAAAUGCAAGC ((....-((((((((.(.((..-.....))).......(((((..........)))))..))))))))..((((((---(((..((....))....((....)).))))....))))))) ( -24.50) >DroGri_CAF1 55310 96 - 1 GCUAAGCGAGUUAUGAAUGCAAUUUC-GAGUUGGUUAUGAAAAUGAAAACGAUUUU-CCAUGGCAUUCAAUUGCAGCAUUGCU-GCAAA--------------CAGCGGCAAA------- (((..((((....(((((((......-((((((.((((....))))...)))))).-.....))))))).))))))).(((((-((...--------------..))))))).------- ( -31.40) >DroSec_CAF1 33495 115 - 1 GCAAUA-GAGAGCCGAGACCAA-UUCAAGGCGAUUUAUGAAAAUGAAAACAUCUUUUCCACGGCUUUCAAUUGCAA---UGCACCCAAUAGGAAAAGCAAAGGCAACGGCAAAUGCAAGC (((((.-((((((((.(.((..-.....)))..(((((....))))).............)))))))).)))))..---((((.((....))....((...(....).))...))))... ( -31.80) >DroEre_CAF1 29668 116 - 1 GCAAAAUGAGAGCCGAGACCAA-UUCCAGGCGAUUUAUGAAAAUGAAAACAUAUUUUCCGCGGCUUUCAAUUGCAC---UGCUCGCAAUCAGAAGAGCAAAGGCAACGGAAAAUUCAACC ((((..(((((((((.(.((..-.....))).......(((((((......)))))))..))))))))).))))..---(((((..........)))))..(....)............. ( -28.60) >consensus GCAAAA_GAGAGCCGAGACCAA_UUCAAGGCGAUUUAUGAAAAUGAAAACAUCUUUUCCACGGCUUUCAAUUGCAC___UGCACCCAAAAGGAAAAGCAAAGGCAACGGCAAAUGCAAGC ((((...((((((((((......))).......(((((....)))))..............)))))))..)))).............................................. (-12.05 = -12.55 + 0.50)


| Location | 21,162,578 – 21,162,669 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 72.82 |
| Mean single sequence MFE | -21.42 |
| Consensus MFE | -12.26 |
| Energy contribution | -13.20 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.57 |
| SVM decision value | 3.13 |
| SVM RNA-class probability | 0.998511 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
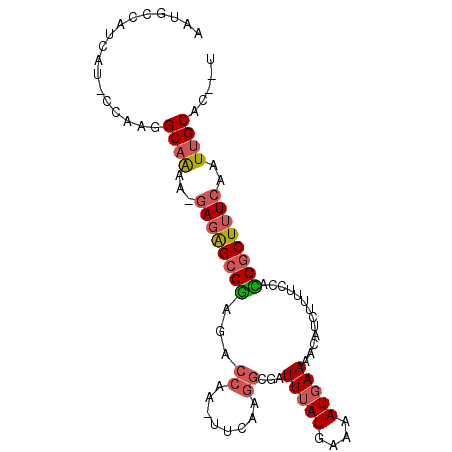
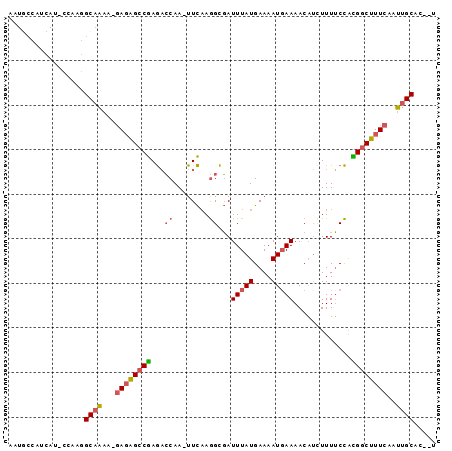
>3L_DroMel_CAF1 21162578 91 - 23771897 AAUACCAUCAU-CCAAGGCAAAA-GAGAGCCGAGACCAA-UUCAAGGCGAUUUAUGAAAAUUAAAACAUCUUUUCCAUGGCUUUCAAUUGCAC--U ...........-.....((((..-((((((((.(.((..-.....))).......(((((..........)))))..))))))))..))))..--. ( -16.70) >DroSec_CAF1 33535 91 - 1 AAUGCCAUCAU-CCAAGGCAAUA-GAGAGCCGAGACCAA-UUCAAGGCGAUUUAUGAAAAUGAAAACAUCUUUUCCACGGCUUUCAAUUGCAA--U ...........-.....(((((.-((((((((.(.((..-.....)))..(((((....))))).............)))))))).)))))..--. ( -23.20) >DroEre_CAF1 29708 92 - 1 AAUGCCAUCAU-CCAAGGCAAAAUGAGAGCCGAGACCAA-UUCCAGGCGAUUUAUGAAAAUGAAAACAUAUUUUCCGCGGCUUUCAAUUGCAC--U ..((((.....-....))))...(((((((((.(.((..-.....))).......(((((((......)))))))..))))))))).......--. ( -22.20) >DroMoj_CAF1 58844 84 - 1 -----------AUUGCAGCUGAGCAAUGGGCAAAUGCAAUUUUAAAGUCAUUUAUGAAAAUGAAAACCAUUUUGCUAUGGCUUUCAAUUGCAGCA- -----------..(((.(((.(....).)))...(((((((..((((((((....(((((((.....)))))).).)))))))).))))))))))- ( -23.60) >consensus AAUGCCAUCAU_CCAAGGCAAAA_GAGAGCCGAGACCAA_UUCAAGGCGAUUUAUGAAAAUGAAAACAUCUUUUCCACGGCUUUCAAUUGCAC__U .................((((...((((((((...((........))...(((((....))))).............))))))))..))))..... (-12.26 = -13.20 + 0.94)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:04:21 2006