
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,127,485 – 21,127,586 |
| Length | 101 |
| Max. P | 0.983897 |

| Location | 21,127,485 – 21,127,586 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 94.65 |
| Mean single sequence MFE | -28.84 |
| Consensus MFE | -22.87 |
| Energy contribution | -23.47 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.84 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.983897 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21127485 101 + 23771897 CGCGGCUCUCUGCCAAAUGGCAGAGUGAAAACUAUUCGCUUUGCUUACGAAUUAGAGCACAAAACAGAAAGCCAACCAAAACAGAGCGGCAAACCGAAAGG .((.((((((((((....))))))(((((.....)))))..(((((........)))))........................)))).))...((....)) ( -31.70) >DroSec_CAF1 178197 101 + 1 CGCGACUCUCUGCCAAAUGGCAGAGUGAAAACUAUUGGCUUUGCUUACGAAUUAGAGCACAAAACAGAAAGCCAACCAAAACAGAGCGGCAAGCCGAAAGG (((..(.(((((((....))))))).).......((((((((((((........)))).(......)))))))))..........))).....((....)) ( -32.00) >DroSim_CAF1 89889 101 + 1 CGCGGCUCUCUGACAAAUGGCAGAGUGAAAACUAUUGGCUUUGCUUACGAAUUAGAGCACAAAACAGAAAGCCAACCAAAACAGAGCGGCAAGCCGAAAGG .((.((((((((.(....).)))(((....))).((((((((((((........)))).(......))))))))).......))))).))...((....)) ( -29.20) >DroEre_CAF1 100595 101 + 1 CGCGGCUCUCUCACAAAUGCCAGAGUGAAAACUAUUCGCUUUGCUUACGAAUUAGAGCACAAAACAGAAAGCCAACCAAAACAGAGCGGCAAGCCGAAAGA .((.(((((.........((.((((((((.....))))))))((((........))))............))..........))))).))....(....). ( -23.01) >DroYak_CAF1 71548 100 + 1 CGCGGCUCUCUCAGCGCUGGCAGAGU-AAAACUAUUCGCUUUGCUUACGAAUUAGAGCACAAAACAGAAAGCCAACCAAAACAGAGCGGCAAGCCGAAAGG .((.(((((........((((.((((-(....)))))((((((.........))))))............))))........))))).))...((....)) ( -28.29) >consensus CGCGGCUCUCUGACAAAUGGCAGAGUGAAAACUAUUCGCUUUGCUUACGAAUUAGAGCACAAAACAGAAAGCCAACCAAAACAGAGCGGCAAGCCGAAAGG .((.(((((........(((((((((.((.....)).)))))((((........))))............))))........))))).))...((....)) (-22.87 = -23.47 + 0.60)

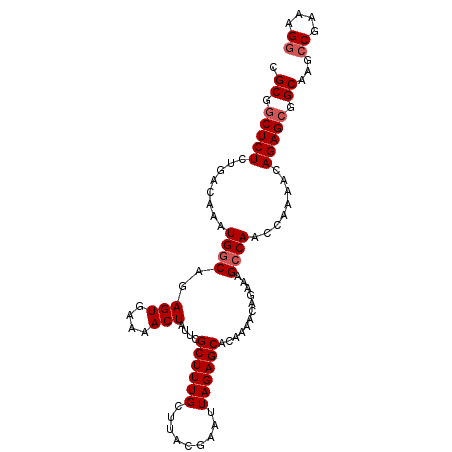
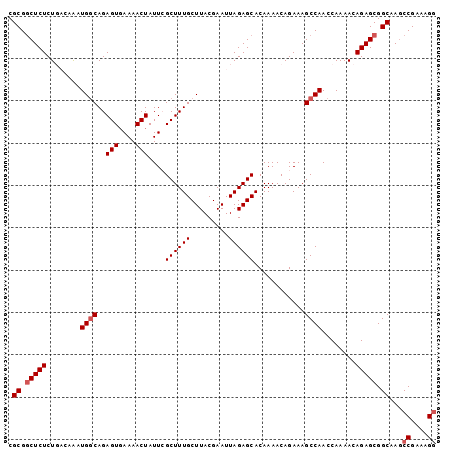
| Location | 21,127,485 – 21,127,586 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 94.65 |
| Mean single sequence MFE | -27.76 |
| Consensus MFE | -25.38 |
| Energy contribution | -25.90 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.09 |
| Structure conservation index | 0.91 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516395 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21127485 101 - 23771897 CCUUUCGGUUUGCCGCUCUGUUUUGGUUGGCUUUCUGUUUUGUGCUCUAAUUCGUAAGCAAAGCGAAUAGUUUUCACUCUGCCAUUUGGCAGAGAGCCGCG .....(((((.((((..(......)..)))).(((.(((((((...(......)...)))))))))).........(((((((....)))))))))))).. ( -30.10) >DroSec_CAF1 178197 101 - 1 CCUUUCGGCUUGCCGCUCUGUUUUGGUUGGCUUUCUGUUUUGUGCUCUAAUUCGUAAGCAAAGCCAAUAGUUUUCACUCUGCCAUUUGGCAGAGAGUCGCG ...........((.((((.......(((((((((((.....((......)).....)).))))))))).........((((((....)))))))))).)). ( -29.30) >DroSim_CAF1 89889 101 - 1 CCUUUCGGCUUGCCGCUCUGUUUUGGUUGGCUUUCUGUUUUGUGCUCUAAUUCGUAAGCAAAGCCAAUAGUUUUCACUCUGCCAUUUGUCAGAGAGCCGCG .....(((((...............(((((((((((.....((......)).....)).)))))))))........(((((.(....).)))))))))).. ( -24.80) >DroEre_CAF1 100595 101 - 1 UCUUUCGGCUUGCCGCUCUGUUUUGGUUGGCUUUCUGUUUUGUGCUCUAAUUCGUAAGCAAAGCGAAUAGUUUUCACUCUGGCAUUUGUGAGAGAGCCGCG .....(((((.((((..(......)..)))).(((.(((((((...(......)...)))))))))).........((((.((....)).))))))))).. ( -26.60) >DroYak_CAF1 71548 100 - 1 CCUUUCGGCUUGCCGCUCUGUUUUGGUUGGCUUUCUGUUUUGUGCUCUAAUUCGUAAGCAAAGCGAAUAGUUUU-ACUCUGCCAGCGCUGAGAGAGCCGCG .....((((((....(((.((.(((((.(((((((.(((((((...(......)...)))))))))).))))..-.....))))).)).))).)))))).. ( -28.00) >consensus CCUUUCGGCUUGCCGCUCUGUUUUGGUUGGCUUUCUGUUUUGUGCUCUAAUUCGUAAGCAAAGCGAAUAGUUUUCACUCUGCCAUUUGUCAGAGAGCCGCG .....(((((.((((..(......)..)))).....(((((((...(......)...)))))))............(((((((....)))))))))))).. (-25.38 = -25.90 + 0.52)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:03:59 2006