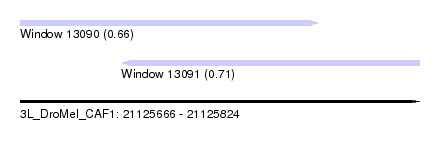

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,125,666 – 21,125,824 |
| Length | 158 |
| Max. P | 0.709528 |

| Location | 21,125,666 – 21,125,784 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.67 |
| Mean single sequence MFE | -30.66 |
| Consensus MFE | -28.58 |
| Energy contribution | -28.94 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.663107 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
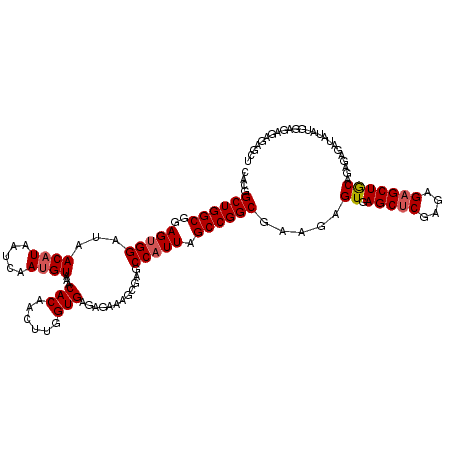
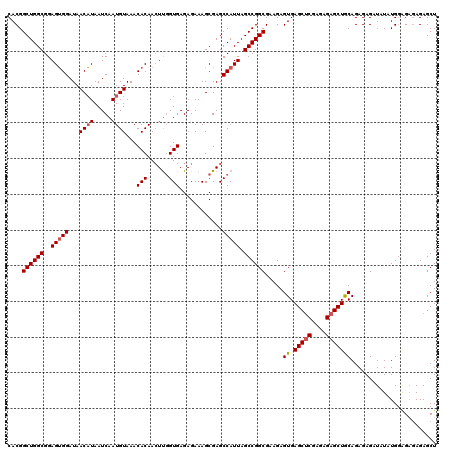
>3L_DroMel_CAF1 21125666 118 + 23771897 CACGGCUGGCGGAGGGGAUAACUUAAUCAAUGUAAACACAACUUUGUGAGAGAAAGCGAGCCAUUAGCCGGCGAAGAGUGAGCUCGAGAGAGCUGCAGAGAGAUAUAUGCAG--AGAGCU (((.((((((....(((((......)))........((((....))))............))....)))))).....)))(((((.......(((((..........)))))--.))))) ( -30.30) >DroSec_CAF1 176388 120 + 1 CACGGCUGGCGGAGUGGAUAACAUAAUCAAUGUAAACACAACUUGGUGAGAGAAAGCGAGCCAUUAGCCGGCGAAGAGUGAGCACGAGAGAGCUGCAGAGAGAUAUAUGGAGAGAGAGCU ....((((((..(((((...((((.....))))........((((.(.......).))))))))).)))))).....((.(((.(....).)))))........................ ( -26.50) >DroSim_CAF1 88051 120 + 1 CACGGCUGGCGGAGUGGAUAACAUAAUCAAUGUAAACACAACUUGGUGAGAGAAAGCGAGCCAUUAGCCGGCGAAGAGUGAGCUCGAGAGAGCUGCAGAGAGAUAUAUGGAGAGAGAGCU ....((((((..(((((...((((.....))))........((((.(.......).))))))))).)))))).....((.(((((....)))))))........................ ( -31.40) >DroEre_CAF1 98795 120 + 1 CACGGCUGGCGGAGUGGAUAACAUAAUCAAUGUAAACACAAGUUGGUGAGAGAAAGCGAGCCAUUAGCCGGCGAAGAGUGAGCUCGAGAGAGCUACAGAGAGAUGUAUGGGGAGAGAGCU (((.((((((..(((((...((((.....))))........(((..(....)..)))...))))).)))))).....)))(((((....)))))(((......))).............. ( -31.60) >DroYak_CAF1 69737 120 + 1 CACGGCUGGCGGAGUGGAUAACAUAAUCAAUGUAAACACAACUCGGUGAGAGAAAGCGAGCCAUUAGCCGGCGAAGAGUGAGCUCGAGAGAGCUACAGAGAGAUAUAUGGGGAGAGAGCU ....((((((..(((((...((((.....))))........((((.(.......).))))))))).)))))).....((.(((((....)))))))........................ ( -33.50) >consensus CACGGCUGGCGGAGUGGAUAACAUAAUCAAUGUAAACACAACUUGGUGAGAGAAAGCGAGCCAUUAGCCGGCGAAGAGUGAGCUCGAGAGAGCUGCAGAGAGAUAUAUGGAGAGAGAGCU ....((((((..(((((...((((.....))))...(((......)))............))))).)))))).....((.(((((....)))))))........................ (-28.58 = -28.94 + 0.36)

| Location | 21,125,706 – 21,125,824 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 91.96 |
| Mean single sequence MFE | -24.76 |
| Consensus MFE | -18.83 |
| Energy contribution | -19.27 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.709528 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
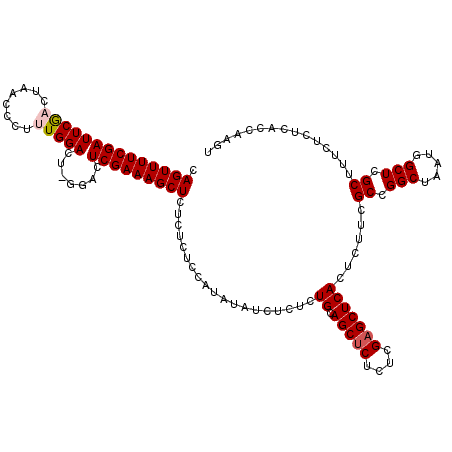
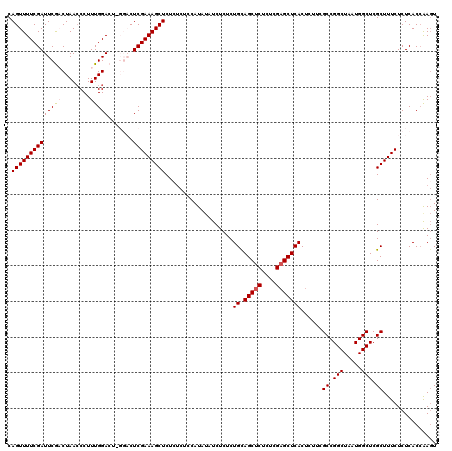
>3L_DroMel_CAF1 21125706 118 - 23771897 CAGUUUUCGAUUCGACUAACCCUUUGGACUGGGACUCGAAAGCUCU--CUGCAUAUAUCUCUCUGCAGCUCUCUCGAGCUCACUCUUCGCCGGCUAAUGGCUCGCUUUCUCUCACAAAGU .((((........))))....(((((((.((((.(((((.((....--(((((..........)))))..)).))))))))).))...((.(((.....))).)).........))))). ( -27.60) >DroSec_CAF1 176428 120 - 1 CAGUUUUCGAUUCGCCUAAACCUUUGGACUCGGACUCGAAAGCUCUCUCUCCAUAUAUCUCUCUGCAGCUCUCUCGUGCUCACUCUUCGCCGGCUAAUGGCUCGCUUUCUCUCACCAAGU .(((((((((((((.((((....))))...)))).)))))))))....................(((((........)))........((((.....))))..))............... ( -21.30) >DroSim_CAF1 88091 120 - 1 CAGUUUUCGAUUCGCCUAACCCUUUGGACUCGGACUCGAAAGCUCUCUCUCCAUAUAUCUCUCUGCAGCUCUCUCGAGCUCACUCUUCGCCGGCUAAUGGCUCGCUUUCUCUCACCAAGU .(((((((((((((.((((....))))...)))).)))))))))....................(((((((....)))))........((((.....))))..))............... ( -25.10) >DroEre_CAF1 98835 114 - 1 CAGUUUUCGAUUCAACUAACCCUUUGGA------CUCGAAAGCUCUCUCCCCAUACAUCUCUCUGUAGCUCUCUCGAGCUCACUCUUCGCCGGCUAAUGGCUCGCUUUCUCUCACCAACU .((((((((((((((........)))))------.)))))))))....................(((((((....))))).)).....((.(((.....))).))............... ( -24.00) >DroYak_CAF1 69777 114 - 1 CAGUUUUCGAUUCGACUAACCCUUUGGA------CUCGAAAGCUCUCUCCCCAUAUAUCUCUCUGUAGCUCUCUCGAGCUCACUCUUCGCCGGCUAAUGGCUCGCUUUCUCUCACCGAGU .((((((((((((((........)))))------.)))))))))................(((.(((((((....))))).)).....((.(((.....))).))...........))). ( -25.80) >consensus CAGUUUUCGAUUCGACUAACCCUUUGGACU_GGACUCGAAAGCUCUCUCUCCAUAUAUCUCUCUGCAGCUCUCUCGAGCUCACUCUUCGCCGGCUAAUGGCUCGCUUUCUCUCACCAAGU .((((((((((((((........))))).......)))))))))...................((.(((((....)))))))......((.(((.....))).))............... (-18.83 = -19.27 + 0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:03:53 2006