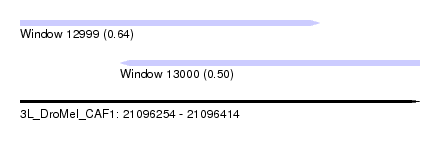
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,096,254 – 21,096,414 |
| Length | 160 |
| Max. P | 0.644620 |

| Location | 21,096,254 – 21,096,374 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.75 |
| Mean single sequence MFE | -33.28 |
| Consensus MFE | -29.65 |
| Energy contribution | -29.90 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.644620 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
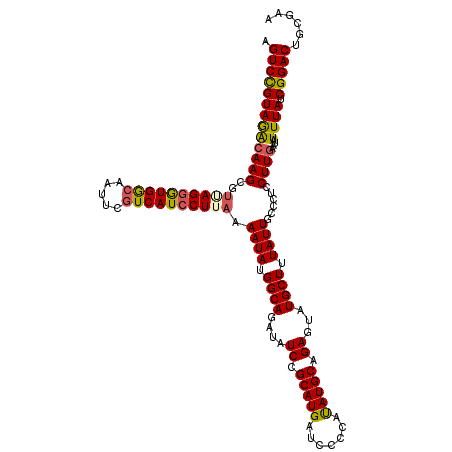
>3L_DroMel_CAF1 21096254 120 + 23771897 UGUCUGUAGAGAAGCGCUAGGGUGACAAUUCGUCAUCCUUAAAAUAUGGCAGAUAUCCGCAUGAUCCCCAUAUGCAGAGUAUGCUUUAUUGCCCUCCUUGAAUAUUUAUCGGACUGCGAA .(((((..(((.((.((.((((((((.....))))))))...((((.((((....((.(((((.......))))).))...)))).)))))).)).)))..........)))))...... ( -30.60) >DroSec_CAF1 146450 120 + 1 AGUCCGUAGGCAAGCGUUAGGGUGGAAAUUCGUCAUCCUUAAAAUAUGGCAGAUAUCCGCAUGAUCCCCAUAUGCAGAGUAUGCUUUAUUGCCCUCCUUGAAUAUUUAUCGGACUGCGAA ((((((((..((((.(..((((((((.((..(((((.........)))))..)).)))(((((.((..(....)..)).)))))......))))))))))..)).....))))))..... ( -31.20) >DroEre_CAF1 68841 120 + 1 AGUCCGUAAACAAGCGUAAGGGUGGCAAUUCGUCAUCCUUAAAAUAUGGCAGAUUUCCGCAUGAUCCCCAUAUGCAGAGUAUGCUUUAUUGCCCUCCUUGAAUAUUUAUCGGACUGCGAA ((((((((((((((..((((((((((.....)))))))))).((((.((((...(((.(((((.......))))).)))..)))).))))......))))....)))).))))))..... ( -36.10) >DroYak_CAF1 40674 120 + 1 AGUCCGUAGGCAAGCGUGAGGAUGGCAAUUCGUCAUACUUAAAAUAUGGCAGAUAUCCGCAUGAUCCCCACAUGCAGAGCAUGCUUUAUUGCCCUCCUUGAAUAUUUAUCGGACUGCGAA ((((((..........((((((.((((((..((((((.......))))))........(((((.......)))))((((....)))))))))).)))))).........))))))..... ( -35.21) >consensus AGUCCGUAGACAAGCGUUAGGGUGGCAAUUCGUCAUCCUUAAAAUAUGGCAGAUAUCCGCAUGAUCCCCAUAUGCAGAGUAUGCUUUAUUGCCCUCCUUGAAUAUUUAUCGGACUGCGAA .(((((((((((((..((((((((((.....)))))))))).((((.((((....((.(((((.......))))).))...)))).))))......))))....)))).)))))...... (-29.65 = -29.90 + 0.25)


| Location | 21,096,294 – 21,096,414 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.54 |
| Mean single sequence MFE | -26.03 |
| Consensus MFE | -24.48 |
| Energy contribution | -24.42 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.94 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21096294 120 - 23771897 GACCUAUAACACAUGUACAUAUACUCGUAUAUAGUAUAUAUUCGCAGUCCGAUAAAUAUUCAAGGAGGGCAAUAAAGCAUACUCUGCAUAUGGGGAUCAUGCGGAUAUCUGCCAUAUUUU ............((((((........)))))).(((.(((((((((((((((.......))..((((.((......))...))))........))))..))))))))).)))........ ( -26.50) >DroSec_CAF1 146490 120 - 1 GACCUAUAACAAAUGUACAUAUACUCGUAUAUAGUAUAUAUUCGCAGUCCGAUAAAUAUUCAAGGAGGGCAAUAAAGCAUACUCUGCAUAUGGGGAUCAUGCGGAUAUCUGCCAUAUUUU ............((((((........)))))).(((.(((((((((((((((.......))..((((.((......))...))))........))))..))))))))).)))........ ( -26.50) >DroEre_CAF1 68881 114 - 1 GACCUAUAACACAUGUACAUAU------AUAUUGUAUAUAUUCGCAGUCCGAUAAAUAUUCAAGGAGGGCAAUAAAGCAUACUCUGCAUAUGGGGAUCAUGCGGAAAUCUGCCAUAUUUU ..((((((....(((((((...------....)))))))....(((((((.............)))..((......)).....)))).))))))......((((....))))........ ( -24.12) >DroYak_CAF1 40714 114 - 1 GACCUAUAACACAUGUACAUAU------AUAUAGUAUAUAUUCGCAGUCCGAUAAAUAUUCAAGGAGGGCAAUAAAGCAUGCUCUGCAUGUGGGGAUCAUGCGGAUAUCUGCCAUAUUUU ......................------((((.(((.(((((((((((((((.......))..(.((((((........)))))).)......))))..))))))))).))).))))... ( -27.00) >consensus GACCUAUAACACAUGUACAUAU______AUAUAGUAUAUAUUCGCAGUCCGAUAAAUAUUCAAGGAGGGCAAUAAAGCAUACUCUGCAUAUGGGGAUCAUGCGGAUAUCUGCCAUAUUUU ..((((((....(((((((((((....))))).))))))....(((((((.............)))..((......)).....)))).))))))......((((....))))........ (-24.48 = -24.42 + -0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:02:20 2006