
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,082,973 – 21,083,160 |
| Length | 187 |
| Max. P | 0.749389 |

| Location | 21,082,973 – 21,083,080 |
|---|---|
| Length | 107 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.75 |
| Mean single sequence MFE | -26.10 |
| Consensus MFE | -15.18 |
| Energy contribution | -18.05 |
| Covariance contribution | 2.87 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.749389 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21082973 107 - 23771897 CGCUGCGGCACCGGGAAUUAAACA----------AGCUAUAAAGCAUAUUGAAAUGUAUAUGCAAAGGUAUAUAUAUAUACA-AUAUAUAUCCGCUCUGUUAUCAGCUGAAACAUUAA-- .((((.((((.((((.........----------.(((....)))((((((..((((((((((....))))))))))...))-))))...))))...))))..))))...........-- ( -24.90) >DroSec_CAF1 138297 118 - 1 CGCUGCGGCACCGGGAAUUAAACAUCAAAUGUGUUGCUAUAAAGCAUAUUGAAAUGUAUAUACAAAGGUAUAAACAUAUAU--AUAUAUAUCCGGUCUGUUAUCAGCUGAAACAUAAACA .(((((((.((((((.........((((.((((((.......)))))))))).((((.(((((....))))).))))....--.......)))))))))....))))............. ( -28.60) >DroEre_CAF1 60169 116 - 1 CGCUGCGGCACCGGGAAUUAAAUAUCAAAUAUGUUGCUUUAAAGCAUAUUGAAAUGUACUUACAGAGGCAUAU----ACAUAUGUAUAUAUCCGGCCUGUUAUCAGCCGAAACAUAAACA .((((.(((((((((......((((..((((((((.......))))))))...))))............((((----((....)))))).)))))..))))..))))............. ( -24.20) >DroYak_CAF1 32401 120 - 1 CGCUGCGGCACCGGGAAUUAAACAUCAAAAUUGCUGAUAUAAAGCAUAUUGAAAUGUAUUUACAAAGGUAUUUAUGUAUAUAUAUGUAUAUCCGGUCUGUUAUCAGCUGAAACAUAAACA .(((((((.((((((.........((((...((((.......))))..)))).((((((.((((..........)))).)))))).....)))))))))....))))............. ( -26.70) >consensus CGCUGCGGCACCGGGAAUUAAACAUCAAAU_UGUUGCUAUAAAGCAUAUUGAAAUGUAUAUACAAAGGUAUAUAUAUAUAUA_AUAUAUAUCCGGUCUGUUAUCAGCUGAAACAUAAACA .(((((((.((((((....................(((....)))........((((((((((....)))))))))).............)))))))))....))))............. (-15.18 = -18.05 + 2.87)


| Location | 21,083,050 – 21,083,160 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.69 |
| Mean single sequence MFE | -33.25 |
| Consensus MFE | -28.74 |
| Energy contribution | -28.80 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.86 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21083050 110 + 23771897 AUAGCU----------UGUUUAAUUCCCGGUGCCGCAGCGGAAACGCAAAAUGGCAAGGCAAGUCAACGCGACGACAGUGGAAAAUGGCGCGUUGAUGUCGUUGUCAUUAUUCAAAUUAG .(((.(----------((..((((......(((((..(((....)))....))))).((((((((((((((....((.(....).)).)))))))))....)))))))))..))).))). ( -33.10) >DroSec_CAF1 138375 120 + 1 AUAGCAACACAUUUGAUGUUUAAUUCCCGGUGCCGCAGCGGAAACGCAAAAUGGCAAGGCAAAUCAGCGCGACGACAGUGGAAAAUGGCGCGUUGAUGUCGUUGUCAUUAUUCAAAUUAG ...((..(((....((........))...)))..)).(((....)))..((((((((((((..((((((((....((.(....).)).)))))))))))).))))))))........... ( -36.50) >DroSim_CAF1 44522 120 + 1 AUAGCAACACAUUUGAUGUUUAAUUCCCGGUGCCGCAGCGGAAACGCAAAAUGGCAAGGCAAAUCAGCGCAACGACAGUGGAAAAUGGCGCGUUGAUGUCGUUGUCAUUAUUCAAAUUAG ...((..(((....((........))...)))..)).(((....)))..((((((((((((..(((((((.....((.(....).))..))))))))))).))))))))........... ( -32.60) >DroYak_CAF1 32481 120 + 1 AUAUCAGCAAUUUUGAUGUUUAAUUCCCGGUGCCGCAGCGGAAACGCAAAAUGGCAAAGCAAGUCAGCACGAAGGCAGUGGAAAAUGGCGCGUUGAUGUCUUUGUCAUUAUUCAAAUUAG (((((((.....)))))))........((....))..(((....)))..((((((((((...((((((.(....)..(((........)))))))))..))))))))))........... ( -30.80) >consensus AUAGCAACACAUUUGAUGUUUAAUUCCCGGUGCCGCAGCGGAAACGCAAAAUGGCAAGGCAAAUCAGCGCGACGACAGUGGAAAAUGGCGCGUUGAUGUCGUUGUCAUUAUUCAAAUUAG ...........................((....))..(((....)))..((((((((.(...(((((((((....((........)).)))))))))..).))))))))........... (-28.74 = -28.80 + 0.06)

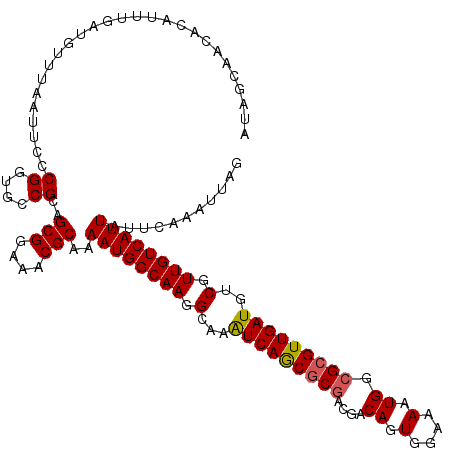

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:01:58 2006