
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,076,720 – 21,076,935 |
| Length | 215 |
| Max. P | 0.995563 |

| Location | 21,076,720 – 21,076,816 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 88.03 |
| Mean single sequence MFE | -32.12 |
| Consensus MFE | -26.10 |
| Energy contribution | -26.35 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.746589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21076720 96 - 23771897 CUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACGAUUGACUGGGUAACUCAGUCUGUGAACC--------------UGCUGAGAGUUCAUGGGA (((.((((((..........(((((((...)))))))((((((((....))))).))).)))))))))((((((((((--------------(....)).))))))))). ( -31.00) >DroSec_CAF1 131889 96 - 1 CUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACAAUUGACUGGGUAACUCAGUCUGUGAACC--------------UGAAGAGAGUUCAUGGGA (((.((((((..........(((((((...)))))))...(((((........))))).)))))))))((((((((((--------------(....)).))))))))). ( -30.80) >DroSim_CAF1 38169 96 - 1 CUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACAAUUGACUGGGUAACUCAGUCUGUGAACC--------------UGAAGAGAGUUCAUGGGA (((.((((((..........(((((((...)))))))...(((((........))))).)))))))))((((((((((--------------(....)).))))))))). ( -30.80) >DroEre_CAF1 54582 110 - 1 CUGCAGUUGCAUAAUUGAAGCGGCGAUUUAAUUGCUGCGGCAGUCUGGCGAUUGACUGGGUAACUCAGGCUGCGAACCGAGCUGAACUGAACUGAUGACAGUUCUUGGGA .(((((((((...((((..((((((((...))))))))..))))...))).....(((((...)))))))))))..((((...((((((.........)))))))))).. ( -35.90) >consensus CUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACAAUUGACUGGGUAACUCAGUCUGUGAACC______________UGAAGAGAGUUCAUGGGA (((.((((((..........(((((((...)))))))...(((((........))))).))))))))).((((((((.......................)))))))).. (-26.10 = -26.35 + 0.25)



| Location | 21,076,738 – 21,076,856 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.64 |
| Mean single sequence MFE | -42.56 |
| Consensus MFE | -38.78 |
| Energy contribution | -38.78 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.91 |
| SVM decision value | 2.59 |
| SVM RNA-class probability | 0.995563 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21076738 118 - 23771897 ACGGGGGAAUCCCGUAUUCGUUUAUGGGGAUUUAUGCCGCCUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACGAUUGACUGGGUAACUCAGUCUGUGAACC-- .((..(((.(((((((......))))))).....((((((..(((((((...(((((......)))))))))))).)))))).)))..))...(((((((...)))))))........-- ( -41.10) >DroSec_CAF1 131907 118 - 1 ACGGGGGAAUCCCGUAUUCGUUUAUGGGGAUUUAUGCCGCCUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACAAUUGACUGGGUAACUCAGUCUGUGAACC-- .((((.((((((((((......))).))))))).((((((..(((((((...(((((......)))))))))))).)))))).)))).((...(((((((...)))))))..))....-- ( -40.40) >DroSim_CAF1 38187 118 - 1 ACGGGGGAAUCCCGUAUUCGUUUACGGGGAUUUAUGCCGCCUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACAAUUGACUGGGUAACUCAGUCUGUGAACC-- (((((.....)))))....(((((((.((((...((((((..(((((((...(((((......)))))))))))).)))))))))).......(((((((...)))))))))))))).-- ( -42.30) >DroEre_CAF1 54612 120 - 1 ACAGGGGAAUCCCGUAUUCGUUCACGGGGAUUUAUGCCGCCUGCAGUUGCAUAAUUGAAGCGGCGAUUUAAUUGCUGCGGCAGUCUGGCGAUUGACUGGGUAACUCAGGCUGCGAACCGA ...((.(((((((((........)).))))))).(((.(((((.((((((.........((((((((...))))))))..(((((........))))).))))))))))).)))..)).. ( -47.60) >DroYak_CAF1 26228 120 - 1 ACGGGGGAAUCCCGUAUUCGUUUACGGGGAUUUAUGCCGCCUGCAGUUGCAUAAUUGGAACGGCGACUUAAUUGCUGCGGCAGUCUGACGAUUGACUGGGCAACUCAGACUGUGAACCAA (((((.....)))))....((((((((((((...((((((..((((((.((....)).)))(....).....))).)))))))))).........((((....).))).))))))))... ( -41.40) >consensus ACGGGGGAAUCCCGUAUUCGUUUACGGGGAUUUAUGCCGCCUGCAGUUGCAUAAUUGGAACGGCGAUUUAAUUGCUGCGGCAGUCUGACGAUUGACUGGGUAACUCAGUCUGUGAACC__ .((((.((((((((((......))).))))))).((((((..(((((((...(((((......)))))))))))).)))))).)))).((...(((((((...)))))))..))...... (-38.78 = -38.78 + 0.00)



| Location | 21,076,776 – 21,076,895 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.21 |
| Mean single sequence MFE | -34.67 |
| Consensus MFE | -29.17 |
| Energy contribution | -29.19 |
| Covariance contribution | 0.02 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.897002 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 21076776 119 + 23771897 CCGCAGCAAUUAAAUCGCCGUUCCAAUUAUGCAACUGCAGGCGGCAUAAAUCCCCAUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGUUGCACCGAAA-A ..((.((.........)).))........(((((((((((.((((((.((((((.............))))))...((((......)))).)))))...).)))))))))))......-. ( -37.32) >DroSec_CAF1 131945 119 + 1 CCGCAGCAAUUAAAUCGCCGUUCCAAUUAUGCAACUGCAGGCGGCAUAAAUCCCCAUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGGCGCACCGAAA-A ..(((((.........((((((.......(((....)))))))))................(((.(((((.....))))).)))(((.((....)).)))))))).((....))....-. ( -34.01) >DroSim_CAF1 38225 119 + 1 CCGCAGCAAUUAAAUCGCCGUUCCAAUUAUGCAACUGCAGGCGGCAUAAAUCCCCGUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGGCGCACCGAAA-A ..(((((.........(((..........(((....))))))(((((.....((((((......))))))......((((......)))).)))))....))))).((....))....-. ( -36.20) >DroEre_CAF1 54652 107 + 1 CCGCAGCAAUUAAAUCGCCGCUUCAAUUAUGCAACUGCAGGCGGCAUAAAUCCCCGUGAACGAAUACGGGAUUCCCCUGUUUUUCUGCGGCAUG-------------GCGCACCGAAAAA ((((((..........((((((.......(((....))))))))).......((((((......))))))..............))))))..((-------------(....)))..... ( -31.11) >DroYak_CAF1 26268 119 + 1 CCGCAGCAAUUAAGUCGCCGUUCCAAUUAUGCAACUGCAGGCGGCAUAAAUCCCCGUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUACAGGCGCACCGAAA-A ((((((.......((((((..........(((....))))))))).......((((((......))))))..............))))))...((((........)))).........-. ( -34.70) >consensus CCGCAGCAAUUAAAUCGCCGUUCCAAUUAUGCAACUGCAGGCGGCAUAAAUCCCCGUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGGCGCACCGAAA_A ((((((..........((((((.......(((....)))))))))................(((.(((((.....))))).)))))))))...((((........))))........... (-29.17 = -29.19 + 0.02)


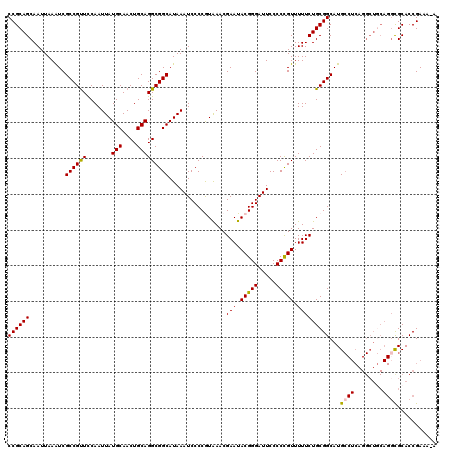
| Location | 21,076,816 – 21,076,935 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.04 |
| Mean single sequence MFE | -32.75 |
| Consensus MFE | -25.18 |
| Energy contribution | -25.60 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.665163 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
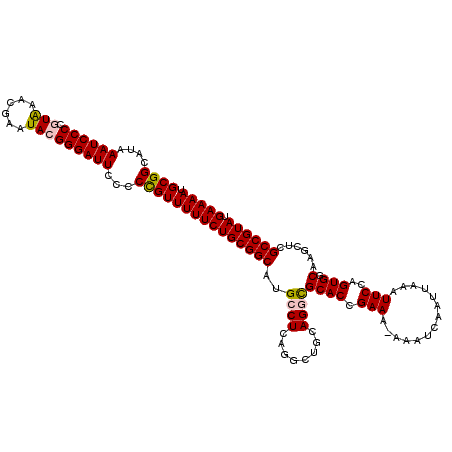
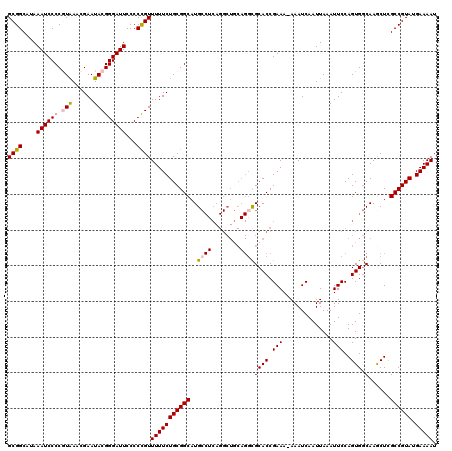
>3L_DroMel_CAF1 21076816 119 + 23771897 GCGGCAUAAAUCCCCAUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGUUGCACCGAAA-AAAUCAAUUAAAUUCCAGUGGCAAGCUCGCCGUAUGAAAAU (((((................(((.(((((.....)))))..)))...(((.((((.(.((.(((....)))))((..-...))............).)))).))).)))))........ ( -33.50) >DroSec_CAF1 131985 119 + 1 GCGGCAUAAAUCCCCAUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGGCGCACCGAAA-AAAUCAAUUAAAUUCCAGUGGCAAGCUCGCCGUAUGAAAAU (((((................(((.(((((.....)))))..)))...(((.((((.(.((.(((....)))))((..-...))............).)))).))).)))))........ ( -33.20) >DroSim_CAF1 38265 119 + 1 GCGGCAUAAAUCCCCGUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGGCGCACCGAAA-AAAUCAAUUAAAUUCCAGUGGCAAGCUCGCCGUAUGAAAAU (((((.........((((...(((.(((((.....)))))..)))))))((.((((.(.((.(((....)))))((..-...))............).)))).))..)))))........ ( -34.70) >DroEre_CAF1 54692 107 + 1 GCGGCAUAAAUCCCCGUGAACGAAUACGGGAUUCCCCUGUUUUUCUGCGGCAUG-------------GCGCACCGAAAAAAAUCAAUUAAAUUCCAGUGCCAAACUUGCCGUAUGAAAAU ((((....((((((((....)).....))))))...))))((((((((((((((-------------((((...(((..............)))..))))))....))))))).))))). ( -30.34) >DroYak_CAF1 26308 119 + 1 GCGGCAUAAAUCCCCGUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUACAGGCGCACCGAAA-AAAUCAAUUAAAUUCCAGUGGCAAACUCGCCGUAUGAAAAU (((((.......((((((......))))))........((((..(((.((....)).)))(((((.((....))((..-...))............)))))))))..)))))........ ( -32.00) >consensus GCGGCAUAAAUCCCCGUAAACGAAUACGGGAUUCCCCCGUUUUUCUGCGGCAUGCCUCAGGCUGCAGGCGCACCGAAA_AAAUCAAUUAAAUUCCAGUGGCAAGCUCGCCGUAUGAAAAU ((((....((((((.(((......)))))))))...))))(((((((((((..((((........))))((((.(((..............)))..))).)......)))))).))))). (-25.18 = -25.60 + 0.42)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:01:42 2006