
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,067,856 – 21,067,976 |
| Length | 120 |
| Max. P | 0.717468 |

| Location | 21,067,856 – 21,067,976 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -39.25 |
| Consensus MFE | -34.33 |
| Energy contribution | -33.95 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.603379 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
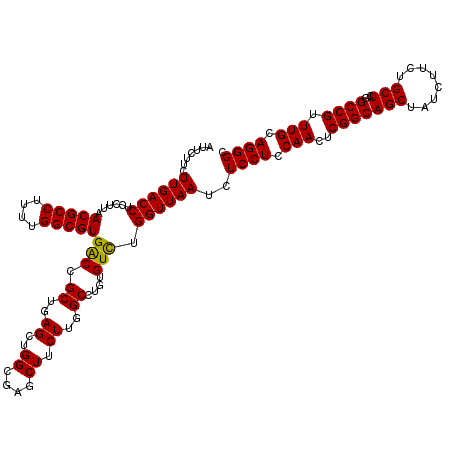
>3L_DroMel_CAF1 21067856 120 + 23771897 AUUCUUCUUGACCUCCUUAACGCCUUUUGGCGUGGCCGCUGAGCUGGCAAGCUUCUUGGCCUGUGUCUGGUUAAUCUCCUCCAACUCGGCAGCUAUCUUCUGCUCGGCCGUUUGAAGGGC .......((((((......(((((....)))))(((((..(((((....)))))..))))).......))))))..((((.(((..(((((((........)))..)))).))).)))). ( -43.30) >DroSec_CAF1 123176 120 + 1 AUUCUUCUUGACCUCCUUAACGCCUUUUGGCGUGACCGCUGAGGAGGCGAGCUUCUUGGCCUGUGUCUGGUUAAUCUCCUCCAACUCGGCAGCUAUCUUCUGCUCGGCCGUUUGCAGGGC ......((((.(((((((((((((....)))))......))))))))))))....((((((.......))))))..((((.(((..(((((((........)))..)))).))).)))). ( -41.70) >DroSim_CAF1 29425 120 + 1 AUUCUUCUUGACCUCCUUAACGCCUUUUGGCGUGACCGCUGAGCUGGCGAGCUUCUUGGCCUGUGUCUGGUUAAUCUCCUCCAACUCGGCAGCUAUCUUCUGCUCGGCCGUUUGCAGGGC .......((((((......(((((....)))))((((((..((..((....)).))..))..).))).))))))..((((.(((..(((((((........)))..)))).))).)))). ( -36.50) >DroYak_CAF1 16259 120 + 1 AUUCUUCUUGACCUCCUUAACGCCUUUUGGCGUGACGGCAGAGCUGGUGAUCUUCUUGGCUUGUGUUUGGUUAAUCUCCUCCAACUCGGCAGCUAUUUUCUGCUCGGCCGUUUGCAGGGC .......((((((.((...(((((....)))))...))..((((..(.(.....))..))))......))))))..((((.(((..(((((((........)))..)))).))).)))). ( -35.50) >consensus AUUCUUCUUGACCUCCUUAACGCCUUUUGGCGUGACCGCUGAGCUGGCGAGCUUCUUGGCCUGUGUCUGGUUAAUCUCCUCCAACUCGGCAGCUAUCUUCUGCUCGGCCGUUUGCAGGGC .......((((((......(((((....)))))(((.((..((..((....)).))..))....))).))))))..((((.(((..(((((((........)))..)))).))).)))). (-34.33 = -33.95 + -0.38)


| Location | 21,067,856 – 21,067,976 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.00 |
| Mean single sequence MFE | -38.45 |
| Consensus MFE | -32.97 |
| Energy contribution | -33.60 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.717468 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
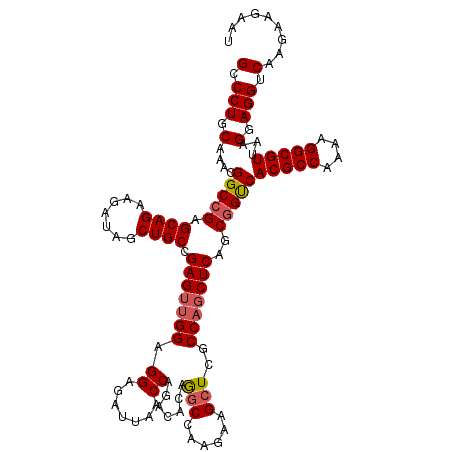
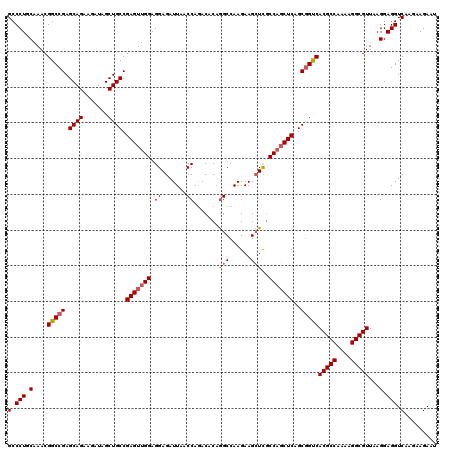
>3L_DroMel_CAF1 21067856 120 - 23771897 GCCCUUCAAACGGCCGAGCAGAAGAUAGCUGCCGAGUUGGAGGAGAUUAACCAGACACAGGCCAAGAAGCUUGCCAGCUCAGCGGCCACGCCAAAAGGCGUUAAGGAGGUCAAGAAGAAU (.(((((....(((((.((((.......)))).(((((((.((.......)).....(((((......))))))))))))..)))))(((((....)))))...))))).)......... ( -45.80) >DroSec_CAF1 123176 120 - 1 GCCCUGCAAACGGCCGAGCAGAAGAUAGCUGCCGAGUUGGAGGAGAUUAACCAGACACAGGCCAAGAAGCUCGCCUCCUCAGCGGUCACGCCAAAAGGCGUUAAGGAGGUCAAGAAGAAU ..(((.(((.((((..(((........)))))))..))).)))................(((......))).(((((((.((((....)(((....)))))).))))))).......... ( -34.40) >DroSim_CAF1 29425 120 - 1 GCCCUGCAAACGGCCGAGCAGAAGAUAGCUGCCGAGUUGGAGGAGAUUAACCAGACACAGGCCAAGAAGCUCGCCAGCUCAGCGGUCACGCCAAAAGGCGUUAAGGAGGUCAAGAAGAAU (.(((.(....(((((.((((.......)))).(((((((..(((.((..((.......))..))....))).)))))))..)))))(((((....)))))...).))).)......... ( -39.30) >DroYak_CAF1 16259 120 - 1 GCCCUGCAAACGGCCGAGCAGAAAAUAGCUGCCGAGUUGGAGGAGAUUAACCAAACACAAGCCAAGAAGAUCACCAGCUCUGCCGUCACGCCAAAAGGCGUUAAGGAGGUCAAGAAGAAU (.(((.(..(((((.(.((((.......)))))(((((((.((.......)).............(.....).))))))).))))).(((((....)))))...).))).)......... ( -34.30) >consensus GCCCUGCAAACGGCCGAGCAGAAGAUAGCUGCCGAGUUGGAGGAGAUUAACCAGACACAGGCCAAGAAGCUCGCCAGCUCAGCGGUCACGCCAAAAGGCGUUAAGGAGGUCAAGAAGAAU (.(((.(....(((((.((((.......)))).(((((((.((.......)).......(((......)))..)))))))..)))))(((((....)))))...).))).)......... (-32.97 = -33.60 + 0.63)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 13:01:21 2006