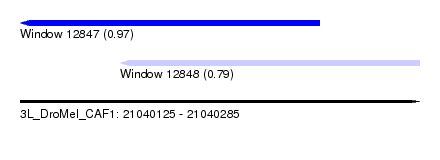

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 21,040,125 – 21,040,285 |
| Length | 160 |
| Max. P | 0.967979 |

| Location | 21,040,125 – 21,040,245 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.81 |
| Mean single sequence MFE | -35.55 |
| Consensus MFE | -34.64 |
| Energy contribution | -34.70 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.97 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.967979 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
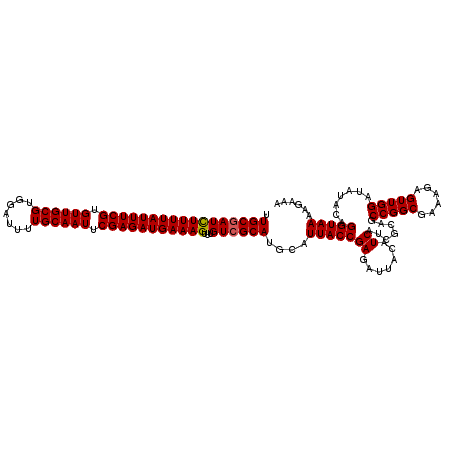
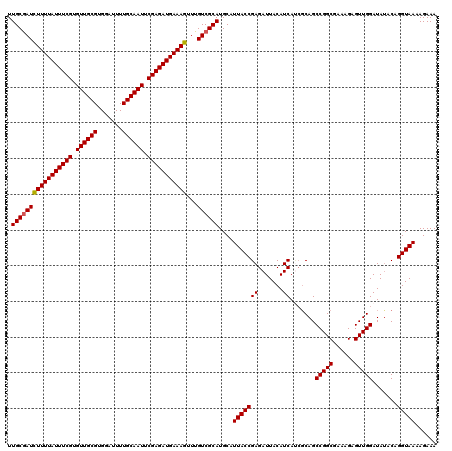
>3L_DroMel_CAF1 21040125 120 - 23771897 UUGCGAUUUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCACCGCAGCCGGCGAAAGAGUUGGAUAUACAGGUAAAAGAAA .(((((((((((((((((.((((((.......)))))).)))))))))))...))))))....(((((((.......)).......(((((......))))).......)))))...... ( -34.60) >DroSec_CAF1 95358 120 - 1 UUGCAAUCUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCAUCGCAGCCGGCGAAAGAGUUGGAUAUACAGGUAAAAGAAA ..((((.(((((((((((.((((((.......)))))).))))))))))).))))........(((((((.......))(((.((((...(....).))))))).....)))))...... ( -33.50) >DroSim_CAF1 1590 120 - 1 UUGCGAUCUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCAUCGCAGCCGGCGAAAGAGUUGGAUACACAGGUAAAAGAAA .(((((((((((((((((.((((((.......)))))).)))))))))))...))))))....(((((((.......))(((.((((...(....).))))))).....)))))...... ( -37.50) >DroEre_CAF1 17411 116 - 1 UUGCGAUCUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCA---CAGCCGGCGAAAGAGUUGGAUAUACAGGUAA-AGAAA .(((((((((((((((((.((((((.......)))))).)))))))))))...))))))....(((((((.......)).---...(((((......))))).......)))))-..... ( -36.60) >consensus UUGCGAUCUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCAUCGCAGCCGGCGAAAGAGUUGGAUAUACAGGUAAAAGAAA .(((((((((((((((((.((((((.......)))))).)))))))))))...))))))....(((((((.......)).......(((((......))))).......)))))...... (-34.64 = -34.70 + 0.06)

| Location | 21,040,165 – 21,040,285 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.89 |
| Mean single sequence MFE | -27.95 |
| Consensus MFE | -27.24 |
| Energy contribution | -27.35 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.787634 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
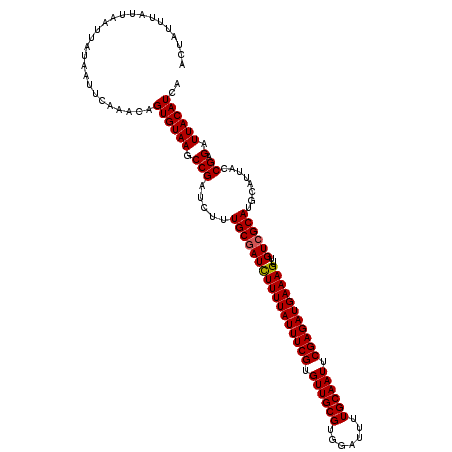
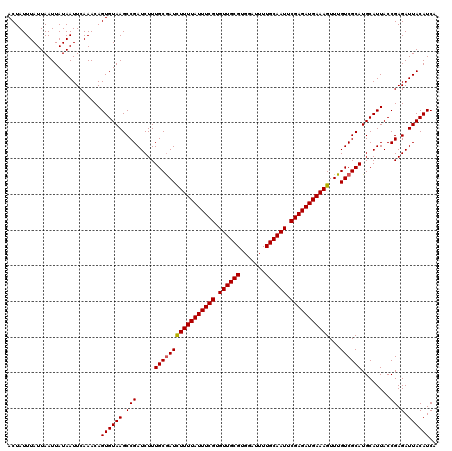
>3L_DroMel_CAF1 21040165 120 - 23771897 ACUAUUUAUUAAUUAUAAUUCAAACAGUGUAAGCCGAUCUUUGCGAUUUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCA ..........................((((((.(((.....(((((((((((((((((.((((((.......)))))).)))))))))))...))))))........)).).)))))).. ( -27.62) >DroSec_CAF1 95398 120 - 1 ACUAUUUAUUAAUUAUAAUUCAAACAGUGUAAGCCGAUCUUUGCAAUCUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCA ...............(((((.....((((((.((........((((.(((((((((((.((((((.......)))))).))))))))))).)))).)).)))))).....)))))..... ( -26.60) >DroSim_CAF1 1630 120 - 1 ACUAUUUAUUAAUUAUAAUUCAAACAGUGUAAGCCGAUCUUUGCGAUCUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCA ..........................((((((.(((.....(((((((((((((((((.((((((.......)))))).)))))))))))...))))))........)).).)))))).. ( -29.62) >consensus ACUAUUUAUUAAUUAUAAUUCAAACAGUGUAAGCCGAUCUUUGCGAUCUUUUAUUUCGUGUUGCGUGGAUUUUGCAAUUCGAGAUGAAAGUUUGUCGCAUGCAUUACCGAGAUUACAUCA ..........................((((((.(((.....(((((((((((((((((.((((((.......)))))).)))))))))))...))))))........)).).)))))).. (-27.24 = -27.35 + 0.11)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:59:43 2006