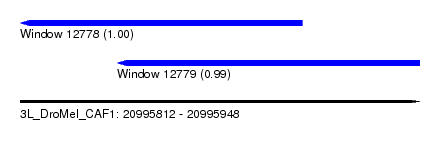

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,995,812 – 20,995,948 |
| Length | 136 |
| Max. P | 0.999975 |

| Location | 20,995,812 – 20,995,908 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 97.29 |
| Mean single sequence MFE | -27.22 |
| Consensus MFE | -26.96 |
| Energy contribution | -26.96 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.27 |
| Structure conservation index | 0.99 |
| SVM decision value | 5.14 |
| SVM RNA-class probability | 0.999975 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20995812 96 - 23771897 UAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCUGCUGAUGUGUUGUGCUGUGUUUAGAACAGUUUC .....((((((((.......))))))))(((((((((..(..(..(((.....)))..)..)..)))))).)))..(((((.......)))))... ( -27.20) >DroSec_CAF1 13411 96 - 1 UAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCUGCUGAUGUGUUGUGCUGUGACUAGAACAGUUCC ....(((((....((((.(((((...(((((((((((..(..(..(((.....)))..)..)..)))))).)))))...))))))))).))))).. ( -27.60) >DroSim_CAF1 20836 96 - 1 UAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCUGCUGAUGUGUUGUGCUGUGACUAGAACAGUUCC ....(((((....((((.(((((...(((((((((((..(..(..(((.....)))..)..)..)))))).)))))...))))))))).))))).. ( -27.60) >DroEre_CAF1 22099 95 - 1 UAAAAGCUGAAUAU-UGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCUGCUGAUGUGUUUUGCUGUGACUAGAACAGUUCC .....((((((((.-.....))))))))(((((((((..(..(..(((.....)))..)..)..)))))).)))..(((((.......)))))... ( -27.10) >DroYak_CAF1 19513 96 - 1 UAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCUGCUGAUGUGUUUUGCUGUGACUAGAACAGUCCC .....((((((((.......))))))))(((((((((..(..(..(((.....)))..)..)..)))))).)))..(((((.......)))))... ( -26.60) >consensus UAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCUGCUGAUGUGUUGUGCUGUGACUAGAACAGUUCC .....((((((((.......))))))))(((((((((..(..(..(((.....)))..)..)..)))))).)))..(((((.......)))))... (-26.96 = -26.96 + 0.00)


| Location | 20,995,845 – 20,995,948 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 95.73 |
| Mean single sequence MFE | -23.76 |
| Consensus MFE | -20.64 |
| Energy contribution | -21.04 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.87 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992328 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20995845 103 - 23771897 ACAUACAGAUCCCAAGCUCUCGAAAAGAACAACCUAAUGCUAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCU .............((((.((.((((((.((((((....(((((..((((((((.......)))))))).....)))))))))))..)))))).))..)))).. ( -25.00) >DroSec_CAF1 13444 103 - 1 AAAUACAGAUCCUAACCGCUCGAAAAGAACAACCUAAUGCUAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCU .............(((.(((.((((((.((((((....(((((..((((((((.......)))))))).....)))))))))))..)))))).))).)))... ( -26.60) >DroSim_CAF1 20869 103 - 1 AAAUACAGAUCCCAAGCGCUCGAAAAGAACAACCUAAUGCUAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCU .............(((((((.((((((.((((((....(((((..((((((((.......)))))))).....)))))))))))..)))))).)).))))).. ( -26.30) >DroEre_CAF1 22132 102 - 1 AAAUACAAAUCACAAGCUCACAAAAAGAACAACCUAAUGCUAAAAGCUGAAUAU-UGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCU ..............((((....(((((.((((((....(((((..((((((((.-.....)))))))).....)))))))))))..)))))..))))...... ( -20.70) >DroYak_CAF1 19546 103 - 1 AAAUACAAAUCCCAAGCUCACAAAAAGAACAACCUAAUGCUAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCU ..............((((....(((((.((((((....(((((..((((((((.......)))))))).....)))))))))))..)))))..))))...... ( -20.20) >consensus AAAUACAGAUCCCAAGCUCUCGAAAAGAACAACCUAAUGCUAAAAGCUGAAUAUUUGUGUUAUUUAGCAACUAUUAGUGGUUGUGGCUUUUCUAGUUGUUUCU .....................((((((.((((((....(((((..((((((((.......)))))))).....)))))))))))..))))))........... (-20.64 = -21.04 + 0.40)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:36 2006