
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,994,161 – 20,994,251 |
| Length | 90 |
| Max. P | 0.773132 |

| Location | 20,994,161 – 20,994,251 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 78.43 |
| Mean single sequence MFE | -27.98 |
| Consensus MFE | -17.09 |
| Energy contribution | -17.97 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.644918 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20994161 90 + 23771897 UUCCACUUCCUCUUCCGUUCGGAGAGAA-ACGGAGAGACUGAGAGAGCGAGACAGAGAG---CAACUGGUGCCGGAGAAGGAGAGGGAGUGUAU---- ...((((((((((((..(((.(......-.(((.....))).....(((...(((....---...))).)))).)))..))))))))))))...---- ( -27.90) >DroSec_CAF1 11808 90 + 1 UUCCAUUUCCUCUUCCGUUCGGAGAGAA-ACGGAGACACUGAGAGAGCGAGAUAGAGAG---CGACUGGUGCCGGAGAAGGAGAGGGAGUGUAU---- ...((((((((((((..(((((......-..(....)(((.((...((..........)---)..))))).)))))...))))))))))))...---- ( -26.50) >DroSim_CAF1 19212 90 + 1 UUCCACUUCCUCUUCCGUUCGGAGAGAA-ACGGAGACACUGAGAGAGCGAGACAGAGAG---CGACUGGUGCCGGAGAAGGAGAGGGAGUGUAU---- ...(((((((((((((((((........-..(....).(((...........))).)))---)).(((....)))....))))))))))))...---- ( -30.30) >DroYak_CAF1 18079 91 + 1 UUCCACUUCCUCUUCCGUUCGGAGAGAAAACGGAGACACUGAGAGGGAGAGACAGAGAG---CGACUGGUGCCGGAGAAAGAGAGGGAGUAUAU---- ....((((((((((.(((((...........(....).((....))..........)))---)).(((....))).....))))))))))....---- ( -26.50) >DroAna_CAF1 10504 82 + 1 UUCCACUUCCUCUUCCGCUCUUAG------------CUCUUA--AAGCCGGAGAGAGAGUUGAGUCUGGCUU--GGAAAAGAGAGAGAGGGUGUGAGA (((((((..((((((((..(((((------------(((((.--..........))))))))))..)))(((--....))).)))))..)))).))). ( -28.70) >consensus UUCCACUUCCUCUUCCGUUCGGAGAGAA_ACGGAGACACUGAGAGAGCGAGACAGAGAG___CGACUGGUGCCGGAGAAGGAGAGGGAGUGUAU____ ...((((((((((((..(((((..............(.(((...........))).)..............)))))...))))))))))))....... (-17.09 = -17.97 + 0.88)



| Location | 20,994,161 – 20,994,251 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 78.43 |
| Mean single sequence MFE | -22.54 |
| Consensus MFE | -11.36 |
| Energy contribution | -11.96 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.29 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.50 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.773132 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20994161 90 - 23771897 ----AUACACUCCCUCUCCUUCUCCGGCACCAGUUG---CUCUCUGUCUCGCUCUCUCAGUCUCUCCGU-UUCUCUCCGAACGGAAGAGGAAGUGGAA ----...((((.(((((........(((..(((...---....)))....)))...........(((((-((......)))))))))))).))))... ( -22.10) >DroSec_CAF1 11808 90 - 1 ----AUACACUCCCUCUCCUUCUCCGGCACCAGUCG---CUCUCUAUCUCGCUCUCUCAGUGUCUCCGU-UUCUCUCCGAACGGAAGAGGAAAUGGAA ----......(((...((((..((((((((.((..(---(..........))...))..)))))...((-((......)))))))..))))...))). ( -21.20) >DroSim_CAF1 19212 90 - 1 ----AUACACUCCCUCUCCUUCUCCGGCACCAGUCG---CUCUCUGUCUCGCUCUCUCAGUGUCUCCGU-UUCUCUCCGAACGGAAGAGGAAGUGGAA ----...((((.(((((........(((((.((..(---(..........))...))..)))))(((((-((......)))))))))))).))))... ( -25.50) >DroYak_CAF1 18079 91 - 1 ----AUAUACUCCCUCUCUUUCUCCGGCACCAGUCG---CUCUCUGUCUCUCCCUCUCAGUGUCUCCGUUUUCUCUCCGAACGGAAGAGGAAGUGGAA ----......(((....(((((((.((((((((...---....))).............)))))(((((((.......))))))).))))))).))). ( -21.71) >DroAna_CAF1 10504 82 - 1 UCUCACACCCUCUCUCUCUUUUCC--AAGCCAGACUCAACUCUCUCUCCGGCUU--UAAGAG------------CUAAGAGCGGAAGAGGAAGUGGAA ((.(((..(((((((((((((((.--(((((.((............)).)))))--...)))------------..))))).)).)))))..))))). ( -22.20) >consensus ____AUACACUCCCUCUCCUUCUCCGGCACCAGUCG___CUCUCUGUCUCGCUCUCUCAGUGUCUCCGU_UUCUCUCCGAACGGAAGAGGAAGUGGAA .......((((.((((((((((..((((....)))).......(((...........)))..................))).)).))))).))))... (-11.36 = -11.96 + 0.60)


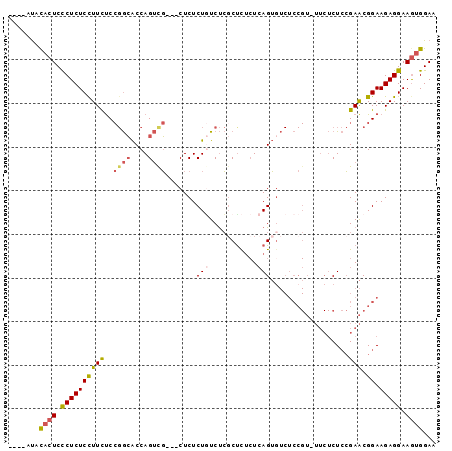
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:58:31 2006