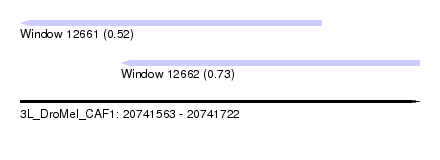
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,741,563 – 20,741,722 |
| Length | 159 |
| Max. P | 0.725845 |

| Location | 20,741,563 – 20,741,683 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.22 |
| Mean single sequence MFE | -42.73 |
| Consensus MFE | -39.43 |
| Energy contribution | -39.77 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.92 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.519107 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20741563 120 - 23771897 AGCUGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUCAAUCCGCAUGUGCCCAAGUAUCUGCUUGAGACAAAGCGACUUAUUCUACCGCCAGAGCACAUUUUGAAGCGCGG .(((((((((.(((((.....)))......((.(((((((((((......(((...(((.(((((....))))).).))..)))....))))))))))).)))).))))))..))).... ( -43.60) >DroSim_CAF1 20097 120 - 1 AGCAGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUCAAUCCGCAUGUGCCCAAGUAUCUGCUGGAGACAAAGCGACUUAUUCUACCGCCAGAGCACAUUUUGAAGCGCGG .((.((((((.(((((.....)))......((.(((((((((((....((((((.((((....)))).)))((....))..).))...))))))))))).)))).))))))...)).... ( -42.40) >DroYak_CAF1 31498 120 - 1 UGCAGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUUAAUCCGCAUGUGCCCAAGUAUCUGCUGGAGACAAAGCGACUUAUUCUACCGCCAGAGCACAUUUUGAAGCGAGG .((.((((((.(((((.....)))......((.(((((((((((....((((((.((((....)))).)))((....))..).))...))))))))))).)))).))))))...)).... ( -42.20) >consensus AGCAGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUCAAUCCGCAUGUGCCCAAGUAUCUGCUGGAGACAAAGCGACUUAUUCUACCGCCAGAGCACAUUUUGAAGCGCGG .((.((((((.(((((.....)))......((.(((((((((((......(((.(((..(((.((....)))))..)))..)))....))))))))))).)))).))))))...)).... (-39.43 = -39.77 + 0.33)

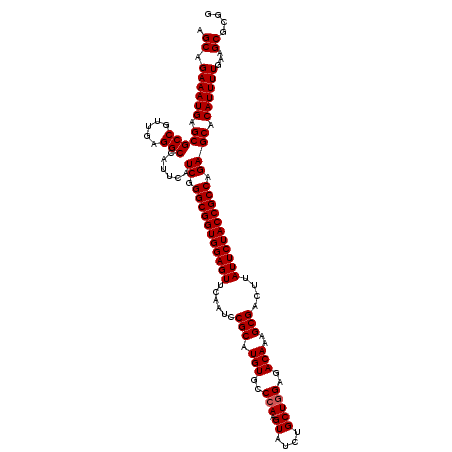
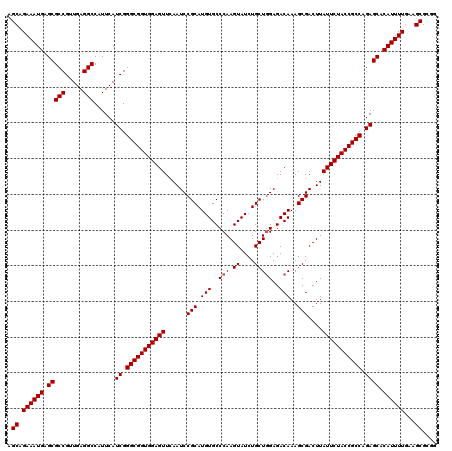
| Location | 20,741,603 – 20,741,722 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 96.64 |
| Mean single sequence MFE | -42.10 |
| Consensus MFE | -41.11 |
| Energy contribution | -41.33 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.725845 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20741603 119 - 23771897 GUUUUCGCCAGACAUAGCAUCCAAGCGAGGUCUAAGUCAAGCUGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUCAAUCCGCAUGUGCCCAAGUAUCUGCUUGAGACAA (((......((((...((......))...))))...((((((.((.(((((.(((.....)))..))))).((((.(((((......)))))....)))).....)).))))))))).. ( -38.30) >DroSim_CAF1 20137 119 - 1 GUUUUCGCCAGACAUAGCAUCCAAGCGAGGUCUAAGUCCAGCAGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUCAAUCCGCAUGUGCCCAAGUAUCUGCUGGAGACAA (((......((((...((......))...))))...(((((((((.(((((.(((.....)))..))))).((((.(((((......)))))....)))).....)))))))))))).. ( -45.30) >DroYak_CAF1 31538 119 - 1 GUUUUCGCCAGACAUUGCAUCCAAGCGAGGUCUAAGUCCUGCAGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUUAAUCCGCAUGUGCCCAAGUAUCUGCUGGAGACAA ((((((...((((.((((......)))).)))).......(((((.(((((.(((.....)))..))))).((((.(((((......)))))....)))).....))))).)))))).. ( -42.70) >consensus GUUUUCGCCAGACAUAGCAUCCAAGCGAGGUCUAAGUCCAGCAGAAAUGAGCGCCGUUGAGGCCAUUCAUCGGGCGGUGGAGUUCAAUCCGCAUGUGCCCAAGUAUCUGCUGGAGACAA (((......((((...((......))...))))...(((((((((.(((((.(((.....)))..))))).((((.(((((......)))))....)))).....)))))))))))).. (-41.11 = -41.33 + 0.22)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:56:42 2006