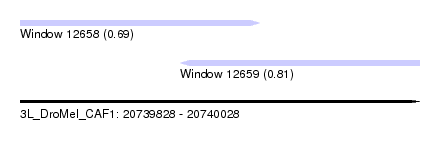
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,739,828 – 20,740,028 |
| Length | 200 |
| Max. P | 0.814884 |

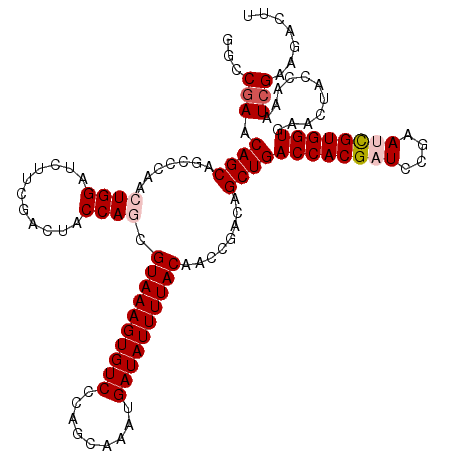
| Location | 20,739,828 – 20,739,948 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.67 |
| Mean single sequence MFE | -31.28 |
| Consensus MFE | -25.54 |
| Energy contribution | -26.38 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.690269 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20739828 120 + 23771897 AACCGAACAGCAGCCCAACUGGAUCUUCGACUACCAGCGUAAAGUGUCCCAGCAAAUGAUAUUUUACAACCGACAGCUGACCACGAUCCGAAUCGUGGUGAACUACCAAAUCGAAGACUU .......(((........)))..(((((((....((((((((((((((.........))))))))))........))))((((((((....))))))))...........)))))))... ( -31.80) >DroSec_CAF1 15308 120 + 1 GGCCGAACAGCAGCCCAACUGGAUCUUCGACUACCAGCGUAAAGUGUCCCAGCAAAUGAUAUUUUACAACCGACAGCUGACCACGAUCCGAAUUGUGGUGAACUACCAAAUCGAAGACUU (((.(.....).)))........(((((((....((((((((((((((.........))))))))))........))))((((((((....))))))))...........)))))))... ( -31.00) >DroSim_CAF1 18406 120 + 1 GGCCGAACAGCAGCCCAACUGGAUCUUCGACUACCAGCGUAAAGUGUCCCAGCAAAUGAUAUUUUACAACCGACAGCUGACCACGAUCCGAAUCGUGGUGAACUACCAAAUCGAAGACUU (((.(.....).)))........(((((((....((((((((((((((.........))))))))))........))))((((((((....))))))))...........)))))))... ( -33.10) >DroEre_CAF1 17053 120 + 1 GGUCGAGCAGCAGCCCAACUGGAUAUUCGACUACCAACGUAAAGUGUCCAAGCAAUUGAUAUUUUACCACCGACAGCUGACCACGUUCCGAAUCGUGGUGAACUACCAAAUCGAAGACUU ..((((.((((...((....))................((((((((((.((....))))))))))))........))))((((((........))))))...........))))...... ( -28.90) >DroYak_CAF1 29720 120 + 1 GGUCGAACAGCAGCCCAACUGGAUAUUCGACUACCAACGUAAAGUGUCGCAGCAAUUGAUAUUUUACCACCGCCAGCUGACCACGUUCCGAAUCGUGGUAAACUACCGAAUGGAAGACUU (((((((.(.(((.....)))..).))))))).(((..(((((((((((.(....))))))))))))...((..((...((((((........))))))...))..))..)))....... ( -31.60) >consensus GGCCGAACAGCAGCCCAACUGGAUCUUCGACUACCAGCGUAAAGUGUCCCAGCAAAUGAUAUUUUACAACCGACAGCUGACCACGAUCCGAAUCGUGGUGAACUACCAAAUCGAAGACUU ...(((.((((.......((((...........)))).((((((((((.........))))))))))........))))((((((((....))))))))...........)))....... (-25.54 = -26.38 + 0.84)


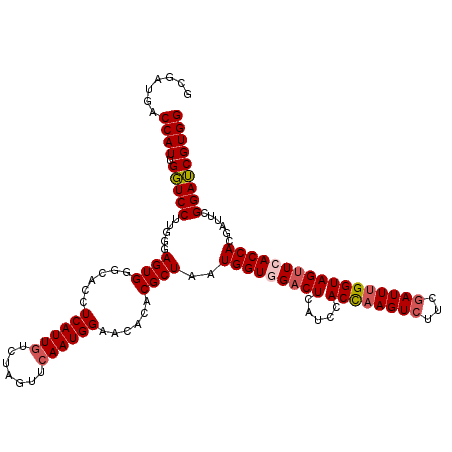
| Location | 20,739,908 – 20,740,028 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.33 |
| Mean single sequence MFE | -39.51 |
| Consensus MFE | -32.32 |
| Energy contribution | -32.88 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.62 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.814884 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20739908 120 - 23771897 GCGAUGACCAUUGGUCCUUGGGAGUGGGCACGUCAUUGUCUAGUUGAAUGGAACACACGCUAAUGGUGGACUCCUCUACUAAGUCUUCGAUUUGGUAGUUCACCACGAUUCGGAUCGUGG ((((((((.....((((........))))..)))))))).....((((.(((...(((.(....))))...))).((((((((((...))))))))))))))(((((((....))))))) ( -38.40) >DroSec_CAF1 15388 120 - 1 GCGAUGACCAUUGGUCCUUGGGAGUGGGCACCUCAUUGUCUAGUUCAAUGGAACACACGCUAAUGGUGGACUCGUCCACCAAGUCUUCGAUUUGGUAGUUCACCACAAUUCGGAUCGUGG .......((((.(((((((((..(((((((......))))..((((....)))).))).))))(((((((((.....((((((((...)))))))))))))))))......))))))))) ( -39.90) >DroSim_CAF1 18486 120 - 1 GCGAUGACCAUUGGUCCUUGGGAGUGGGCACCUCAUUGUCUAGUUCAAUGGAACACACGCUAAUGGUGGACUCGUCCACCAAGUCUUCGAUUUGGUAGUUCACCACGAUUCGGAUCGUGG ......(((((((((((...(((.((((((......)))))).)))...)).......)))))))))(((....)))((((((((...))))))))......(((((((....))))))) ( -40.91) >DroEre_CAF1 17133 120 - 1 GCGAUGACCAUUGUUCCUUUGGAGUGUGCACCUCAUUGUCUAGUUCAAUGGAACACACGCUAAUGGUCGUCUCAUCCACCAAGUCUUCGAUUUGGUAGUUCACCACGAUUCGGAACGUGG (.(((((((((((((((.(((((.((.(((......))).)).))))).)))))).......))))))))).)....((((((((...))))))))......(((((........))))) ( -40.81) >DroYak_CAF1 29800 120 - 1 GCGUUGACCAUUGUUCCUUUGGAGUGGGCACCUCAUUGUCUAAUUCAAUGGAACACACGCUAAUGGUCGACUCAUCCACCAAGUCUUCCAUUCGGUAGUUUACCACGAUUCGGAACGUGG (.(((((((((((((((.(((((.((((((......)))))).))))).)))))).......))))))))).).....(((....((((..(((...........)))...))))..))) ( -37.51) >consensus GCGAUGACCAUUGGUCCUUGGGAGUGGGCACCUCAUUGUCUAGUUCAAUGGAACACACGCUAAUGGUGGACUCAUCCACCAAGUCUUCGAUUUGGUAGUUCACCACGAUUCGGAUCGUGG .......((((.(((((.....((((......((((((.......))))))......))))..(((((((((.....((((((((...)))))))))))))))))......))))))))) (-32.32 = -32.88 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:56:39 2006