| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 2,330,245 – 2,330,352 |
| Length | 107 |
| Max. P | 0.939739 |

| Location | 2,330,245 – 2,330,352 |
|---|---|
| Length | 107 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 79.07 |
| Mean single sequence MFE | -27.17 |
| Consensus MFE | -18.74 |
| Energy contribution | -21.35 |
| Covariance contribution | 2.61 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.939739 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 2330245 107 + 23771897 --GCUUGCGGCACUCAUAAACACAUUUUAUCGAAGGCAGUUAGGCCAAAAAAAAAAACAUAGAAAUGGCCCAAACUUUGGCCAUUAUUCGAACGUGCUGCGACU--GGAAA --..(((((((((...((((.....))))((((((((......))).................(((((((........))))))).)))))..)))))))))..--..... ( -31.50) >DroSec_CAF1 79459 82 + 1 --GCUUGCGGCACUCAUAAACACAUUUUAUCGAAGGCAGUUAGGCC-------------------------AAACUUUGGCCAUUAUUCGAACGUGCUGCGACU--GGAAA --..(((((((((...((((.....))))(((((...(((..((((-------------------------(.....)))))))).)))))..)))))))))..--..... ( -23.60) >DroEre_CAF1 84697 101 + 1 --GAUUGCGGCACUCAUAAACACAUUUUAUCGAAGGCAGUUAGGCCAAA------AACAUAGCGAUGGCCCAAACUUUGGCCAUUAUUCGAACGUGCUGCGACU--GGAGA --..(((((((((...((((.....))))((((((((......)))...------........(((((((........))))))).)))))..)))))))))..--..... ( -32.10) >DroYak_CAF1 82228 101 + 1 --GCUUGCGGCACUCAUAAACACAUUUUAUCGAAGGCAGUUAGGCCAAG------AACAUAGAAAUGGCCCAAACUUUGGCCAUUAUUCGAACGUGCUGCCACU--GAAAA --....(((((((...((((.....))))((((((((......)))...------........(((((((........))))))).)))))..)))))))....--..... ( -29.50) >DroAna_CAF1 77053 102 + 1 --GCUUGCGGCACCUAUAAACACAUUUUAUCGAAGGCA-UUAAGCCAGUAU----AACGAAAAAAUGGCUCAAACUUUGGCCAUUAUUAGAACGUGCUGCGCCU--GCGAA --((.((((((((.....................(((.-....))).....----........(((((((........)))))))........))))))))...--))... ( -27.20) >DroPer_CAF1 103651 102 + 1 AGGCUC---GCUCUCAUAAACACAUUUUAUCGAAAGCAGUUAAGGCGCACA------AAAAAAAAUGGCUAAAACUUUGGCAAUUAUUGGAACGUGCUGCGACUUCGGAAC .((((.---(((.(((((((.....))))).)).)))))))...((((((.------...((.(((.(((((....))))).))).)).....)))).))..(....)... ( -19.10) >consensus __GCUUGCGGCACUCAUAAACACAUUUUAUCGAAGGCAGUUAGGCCAAA______AACAUAGAAAUGGCCCAAACUUUGGCCAUUAUUCGAACGUGCUGCGACU__GGAAA ....(((((((((...((((.....))))((((((((......))).................(((((((........))))))).)))))..)))))))))......... (-18.74 = -21.35 + 2.61)

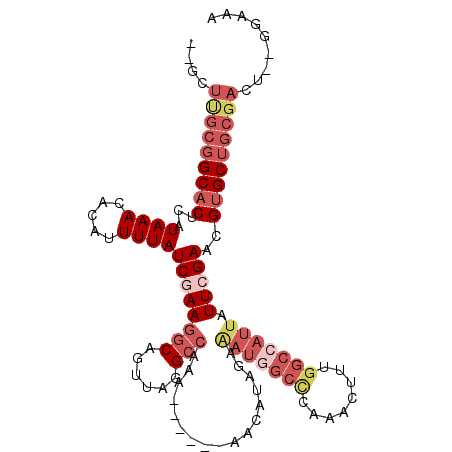
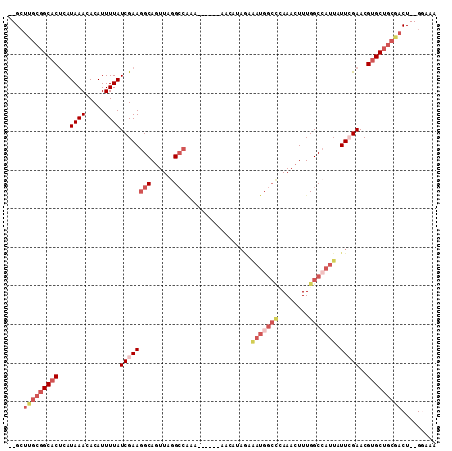
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:50:56 2006