
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,727,512 – 20,727,622 |
| Length | 110 |
| Max. P | 0.957550 |

| Location | 20,727,512 – 20,727,622 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 87.12 |
| Mean single sequence MFE | -27.27 |
| Consensus MFE | -20.40 |
| Energy contribution | -21.27 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.48 |
| SVM RNA-class probability | 0.957550 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
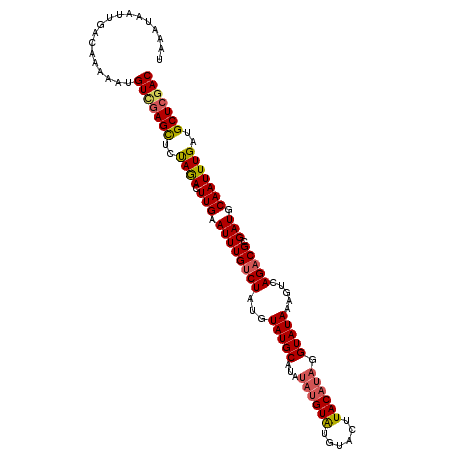
>3L_DroMel_CAF1 20727512 110 + 23771897 UAAAUAAUUGACAAAAAUGUAGAGCUCUAGACUUGAAUUUGUCUAUGUAUGCAUAUAUGUGUGUACUUACAUAGGUAUAAAGUCAGACGCGAUGCAAUUUGAUGCUCGAC .......((((((.(((((((..(((((.(((((.....(..(((((((((((((....))))))..)))))))..)..)))))))).))..))).))))..)).)))). ( -28.60) >DroSec_CAF1 2999 110 + 1 UAAAUAAUUGAUAAAAAUGUCGAGCUCUAGACUUGAAUUUGUCUAUGUAUGCAUAUAUGUAUGUACUUACAUAGGUAUAAAGUCAGACGCGAUGCAAUUUGAUGCUCGAC ..................((((((((((.(((((.....(..(((((((((((((....))))))..)))))))..)..)))))))).((...))........))))))) ( -29.90) >DroSim_CAF1 5349 110 + 1 UAAAUAAUUGACAAAAAUGUCCAGCUCUAGACUUGAAUUUGUCUAUGUAUGCAUAUAUGUAUGUACUUACAUAGGUAUAAAGUCAGACGCGAUGCAAUUUGAUGCUCGAC .........((((....))))..(((((.(((((.....(..(((((((((((((....))))))..)))))))..)..)))))))).))((.(((......)))))... ( -27.50) >DroEre_CAF1 4629 93 + 1 UAAUUAAUUGACAAACAUGUUGAGUUCCAAACUUGAAUUUGCCUCUGUAUGCA-----------------AUAGGUAUAAAGUCAGACGCGAUGCAAUUUGAUGCUCGAC ..................(((((((..((((.(((...((((.(((((((((.-----------------....)))))....)))).))))..)))))))..))))))) ( -23.10) >consensus UAAAUAAUUGACAAAAAUGUCGAGCUCUAGACUUGAAUUUGUCUAUGUAUGCAUAUAUGUAUGUACUUACAUAGGUAUAAAGUCAGACGCGAUGCAAUUUGAUGCUCGAC ..................(((((((..((((.(((.((((((((...(((((...((((((......)))))).))))).....))))).))).)))))))..))))))) (-20.40 = -21.27 + 0.88)


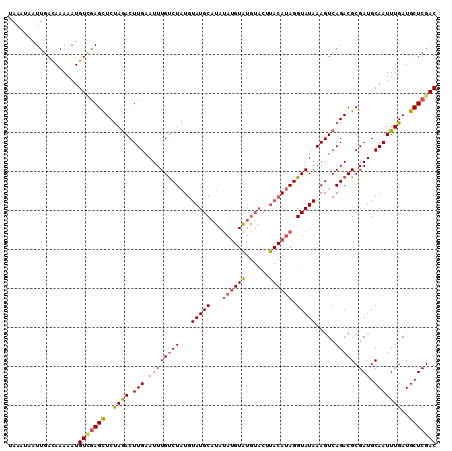
| Location | 20,727,512 – 20,727,622 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 87.12 |
| Mean single sequence MFE | -20.84 |
| Consensus MFE | -13.97 |
| Energy contribution | -16.34 |
| Covariance contribution | 2.37 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.543263 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
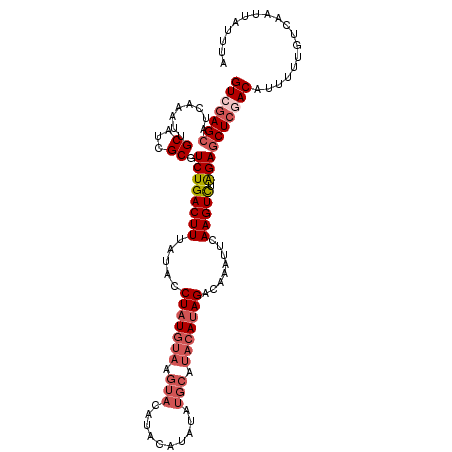
>3L_DroMel_CAF1 20727512 110 - 23771897 GUCGAGCAUCAAAUUGCAUCGCGUCUGACUUUAUACCUAUGUAAGUACACACAUAUAUGCAUACAUAGACAAAUUCAAGUCUAGAGCUCUACAUUUUUGUCAAUUAUUUA ((.((((.((....(((((...((.(((((((((....))).)))).)).))....)))))....(((((........))))))))))).)).................. ( -19.70) >DroSec_CAF1 2999 110 - 1 GUCGAGCAUCAAAUUGCAUCGCGUCUGACUUUAUACCUAUGUAAGUACAUACAUAUAUGCAUACAUAGACAAAUUCAAGUCUAGAGCUCGACAUUUUUAUCAAUUAUUUA (((((((........((...)).((((((((.....(((((((.(((.((....)).))).)))))))........))))).)))))))))).................. ( -26.02) >DroSim_CAF1 5349 110 - 1 GUCGAGCAUCAAAUUGCAUCGCGUCUGACUUUAUACCUAUGUAAGUACAUACAUAUAUGCAUACAUAGACAAAUUCAAGUCUAGAGCUGGACAUUUUUGUCAAUUAUUUA ...(((((......))).))((.((((((((.....(((((((.(((.((....)).))).)))))))........))))).)))))..((((....))))......... ( -23.22) >DroEre_CAF1 4629 93 - 1 GUCGAGCAUCAAAUUGCAUCGCGUCUGACUUUAUACCUAU-----------------UGCAUACAGAGGCAAAUUCAAGUUUGGAACUCAACAUGUUUGUCAAUUAAUUA ((.(((..(((((((.....((.((((.............-----------------......)))).)).......)))))))..))).)).................. ( -14.41) >consensus GUCGAGCAUCAAAUUGCAUCGCGUCUGACUUUAUACCUAUGUAAGUACAUACAUAUAUGCAUACAUAGACAAAUUCAAGUCUAGAGCUCGACAUUUUUGUCAAUUAUUUA (((((((........((...)).((((((((.....(((((((.(((..........))).)))))))........))))).)))))))))).................. (-13.97 = -16.34 + 2.37)

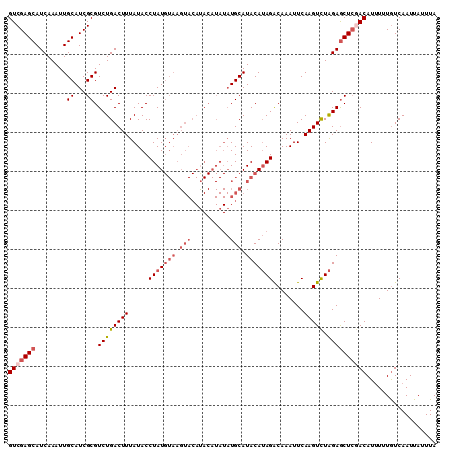
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:56:28 2006