
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,698,243 – 20,698,356 |
| Length | 113 |
| Max. P | 0.877596 |

| Location | 20,698,243 – 20,698,356 |
|---|---|
| Length | 113 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.06 |
| Mean single sequence MFE | -28.85 |
| Consensus MFE | -20.31 |
| Energy contribution | -20.64 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.44 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.861851 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20698243 113 + 23771897 CUGUCGAAAGUGAAUUUACUUACGACAGAAAGUUGCAGCC-------AAUGCCAAUACCAAUGCGAAUGGGAAAACAUCAAAGCCCCCAGGGAACGGCAUCUUUUUGGUAACCAAUUGGC ((((((.(((((....))))).)))))).........(((-------(((.....(((((((((...((((..............))))(....).)))).....)))))....)))))) ( -31.64) >DroEre_CAF1 20142 112 + 1 CUGUCGAAAGUGAAUUUACUUACGACAGAAAGUUGCUGCCAAAUACCAAUACCAAUACCUAUACGAAUGGGAAAACAUCAAAGCCCCCAGGCAACGGCAUCUUUCU--------AUUGGC ..((((.(((((....))))).))))((((((.(((((...................(((((....)))))...........(((....)))..))))).))))))--------...... ( -29.70) >DroYak_CAF1 1094 101 + 1 CUGUCGAAAGUGAAUUUACCUACGACAGAAAGUUUCUGCC-------AAUACCAAUACCAAUACGGAUGGGAAAACAUCAAAGCCCCCAGGCAACGGCAUC----U--------AUUGGC ((((((...(((....)))...)))))).........(((-------((((((............((((......))))...(((....)))...))....----)--------)))))) ( -25.20) >consensus CUGUCGAAAGUGAAUUUACUUACGACAGAAAGUUGCUGCC_______AAUACCAAUACCAAUACGAAUGGGAAAACAUCAAAGCCCCCAGGCAACGGCAUCUUU_U________AUUGGC ((((((.(((((....))))).)))))).........(((...........................((((..............))))(....))))...................... (-20.31 = -20.64 + 0.33)

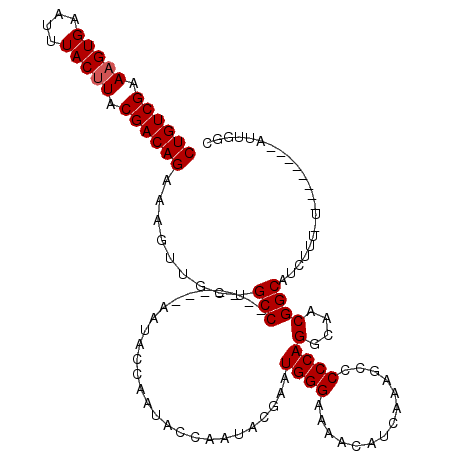

| Location | 20,698,243 – 20,698,356 |
|---|---|
| Length | 113 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.06 |
| Mean single sequence MFE | -33.77 |
| Consensus MFE | -27.77 |
| Energy contribution | -27.67 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.877596 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20698243 113 - 23771897 GCCAAUUGGUUACCAAAAAGAUGCCGUUCCCUGGGGGCUUUGAUGUUUUCCCAUUCGCAUUGGUAUUGGCAUU-------GGCUGCAACUUUCUGUCGUAAGUAAAUUCACUUUCGACAG (((..((((...))))...(((((((((((...))))).(..((((..........))))..)....))))))-------))).........((((((.((((......)))).)))))) ( -31.40) >DroEre_CAF1 20142 112 - 1 GCCAAU--------AGAAAGAUGCCGUUGCCUGGGGGCUUUGAUGUUUUCCCAUUCGUAUAGGUAUUGGUAUUGGUAUUUGGCAGCAACUUUCUGUCGUAAGUAAAUUCACUUUCGACAG (((((.--------..(..(((((((.((((((...((...((((......)))).)).)))))).)))))))..)..))))).........((((((.((((......)))).)))))) ( -35.40) >DroYak_CAF1 1094 101 - 1 GCCAAU--------A----GAUGCCGUUGCCUGGGGGCUUUGAUGUUUUCCCAUCCGUAUUGGUAUUGGUAUU-------GGCAGAAACUUUCUGUCGUAGGUAAAUUCACUUUCGACAG ((((((--------(----(((((((.(((.(((((((......))..)))))...))).)))))))..))))-------))).........((((((.((((......)))).)))))) ( -34.50) >consensus GCCAAU________A_AAAGAUGCCGUUGCCUGGGGGCUUUGAUGUUUUCCCAUUCGUAUUGGUAUUGGUAUU_______GGCAGCAACUUUCUGUCGUAAGUAAAUUCACUUUCGACAG (((................(((((((.(((((((((((......))..)))))........)))).))))))).......))).........((((((.((((......)))).)))))) (-27.77 = -27.67 + -0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:56:12 2006