
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,631,357 – 20,631,469 |
| Length | 112 |
| Max. P | 0.993015 |

| Location | 20,631,357 – 20,631,469 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 85.78 |
| Mean single sequence MFE | -28.85 |
| Consensus MFE | -23.60 |
| Energy contribution | -23.85 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.858672 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20631357 112 + 23771897 AAAUUGUCGAGCAGCCGAUGGUUAUAACCCGAGCCAAACAAGAGCACCCGAAUUCCACCCAUCCAUCCACCCAUCCGCAAU-UUUUUAAGCGGAAAAGGGAUAAAACGGCUGC ..........(((((((.(((((........))))).................................(((.(((((...-.......)))))...)))......))))))) ( -29.30) >DroSec_CAF1 4659 112 + 1 AAAUUGUCGAGCAGCCGAUGGUUAUAACCCGAGCCAAACAAGAGCACCCGAAUUCCACCCAUCCAUCCACCCAUCCGCAAU-UUUUUAAGCGGAAAAGGGAUAAAACGGCUGC ..........(((((((.(((((........))))).................................(((.(((((...-.......)))))...)))......))))))) ( -29.30) >DroSim_CAF1 6271 112 + 1 AAAUUGUCGAGCAGCCGAUGGUUAUAACCCGAGCCAAACAAGAGCACCCGAAUUCCACCCAUCCGUCCACCCAUCCGCAAU-UUUUUAAGCGGAAAAGGGAUAAAACGGCUGC ..........(((((((.(((((........)))))............................((((.....(((((...-.......)))))....))))....))))))) ( -29.60) >DroAna_CAF1 4291 101 + 1 AAA--GUCGAGCAGCCGAUGGCCAUAACCCGAGCCAAUGCAGAGCACCU----------CACCCAUCCGCUCAUGCCCACUUUUUUUAAGUGGAAAAAGGAUGAAAUGGUUCC ...--((((......))))...........((((((.....(((...))----------)...(((((((....))((((((.....)))))).....)))))...)))))). ( -27.20) >consensus AAAUUGUCGAGCAGCCGAUGGUUAUAACCCGAGCCAAACAAGAGCACCCGAAUUCCACCCAUCCAUCCACCCAUCCGCAAU_UUUUUAAGCGGAAAAGGGAUAAAACGGCUGC ..........(((((((.(((((........)))))............................((((.....(((((...........)))))....))))....))))))) (-23.60 = -23.85 + 0.25)

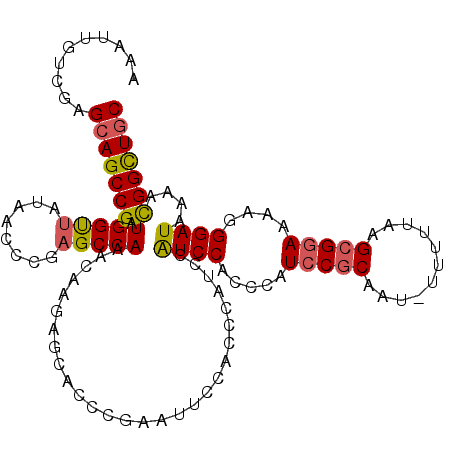
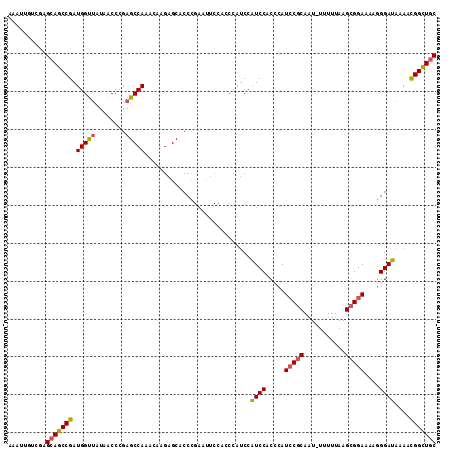
| Location | 20,631,357 – 20,631,469 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 85.78 |
| Mean single sequence MFE | -42.60 |
| Consensus MFE | -33.28 |
| Energy contribution | -35.52 |
| Covariance contribution | 2.25 |
| Combinations/Pair | 1.12 |
| Mean z-score | -3.23 |
| Structure conservation index | 0.78 |
| SVM decision value | 2.37 |
| SVM RNA-class probability | 0.993015 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20631357 112 - 23771897 GCAGCCGUUUUAUCCCUUUUCCGCUUAAAAA-AUUGCGGAUGGGUGGAUGGAUGGGUGGAAUUCGGGUGCUCUUGUUUGGCUCGGGUUAUAACCAUCGGCUGCUCGACAAUUU ((((((....(((((((..(((((.......-...)))))..)).)))))(((((....(((((((((((....))...)))))))))....))))))))))).......... ( -46.40) >DroSec_CAF1 4659 112 - 1 GCAGCCGUUUUAUCCCUUUUCCGCUUAAAAA-AUUGCGGAUGGGUGGAUGGAUGGGUGGAAUUCGGGUGCUCUUGUUUGGCUCGGGUUAUAACCAUCGGCUGCUCGACAAUUU ((((((....(((((((..(((((.......-...)))))..)).)))))(((((....(((((((((((....))...)))))))))....))))))))))).......... ( -46.40) >DroSim_CAF1 6271 112 - 1 GCAGCCGUUUUAUCCCUUUUCCGCUUAAAAA-AUUGCGGAUGGGUGGACGGAUGGGUGGAAUUCGGGUGCUCUUGUUUGGCUCGGGUUAUAACCAUCGGCUGCUCGACAAUUU ((((((((((...(((...(((((.......-...))))).))).)))).(((((....(((((((((((....))...)))))))))....))))))))))).......... ( -45.80) >DroAna_CAF1 4291 101 - 1 GGAACCAUUUCAUCCUUUUUCCACUUAAAAAAAGUGGGCAUGAGCGGAUGGGUG----------AGGUGCUCUGCAUUGGCUCGGGUUAUGGCCAUCGGCUGCUCGAC--UUU (((..((((((((((.....((((((.....))))))((....))....)))))----------))))).)))((....))((((((...((((...)))))))))).--... ( -31.80) >consensus GCAGCCGUUUUAUCCCUUUUCCGCUUAAAAA_AUUGCGGAUGGGUGGAUGGAUGGGUGGAAUUCGGGUGCUCUUGUUUGGCUCGGGUUAUAACCAUCGGCUGCUCGACAAUUU ((((((....(((((((..(((((...........)))))..)).)))))(((((....(((((((((.(........))))))))))....))))))))))).......... (-33.28 = -35.52 + 2.25)

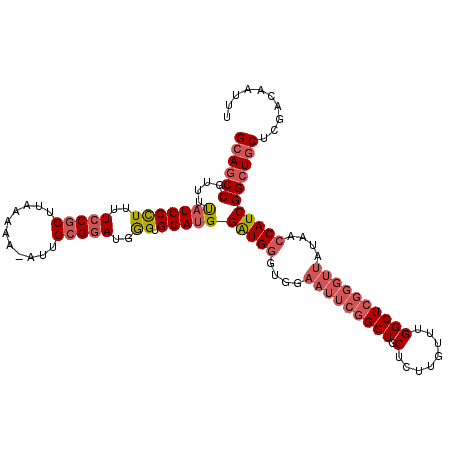
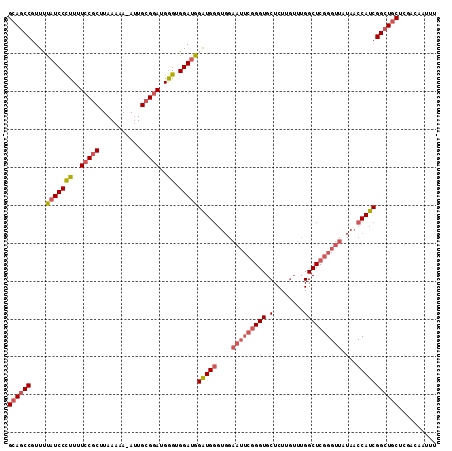
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:55:27 2006