
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,546,623 – 20,546,720 |
| Length | 97 |
| Max. P | 0.510889 |

| Location | 20,546,623 – 20,546,720 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 80.04 |
| Mean single sequence MFE | -34.90 |
| Consensus MFE | -19.21 |
| Energy contribution | -19.72 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.55 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.510889 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20546623 97 - 23771897 ----ACAGA-AACUGUUCCAACUGGCUGCCCC---GCCACUUUAUUACGCUGAU------AAGUUGGCGCCUAAUAACGAUAGUGGAUCGAGGGCCUUCUGGGGCAGCAGG ----((((.-..)))).....(((.(((((((---((((.((((((.....)))------))).))))((((.....((((.....)))).)))).....)))))))))). ( -35.80) >DroPse_CAF1 106026 108 - 1 UGCCACAAAUAACUGUUCCAACUGGUUGCCCCUCUGCCACUUUAUUACGCCAAUGGAAUCAAGUUGGCGCCUAAUAAUAAU---GCUUCGGGGCCGAAGGGGGGCCGAAGG .(((.....(((((((....)).)))))((((((.(((...(((((((((((((........)))))))..))))))....---.(....)))).).))))))))...... ( -33.00) >DroSec_CAF1 88461 97 - 1 ----ACAGA-AACUGUUCCAACUGGCUGCCCC---GUCACUUUAUUACGCUGAU------AAGUUGGCGCCUAAUAACGAUAGUGGAUCGUGGGGCUUCUGGGGCAGCAGG ----((((.-..)))).....(((.(((((((---((((.((((((.....)))------))).))))((((....(((((.....))))).))))....)))))))))). ( -37.90) >DroSim_CAF1 89997 96 - 1 ----ACAGA-AACUGUUCCAACUGGCUGCCCC---GCCACUUUAUUACGCUGAU------AAGUUGGCGCCUAAUAACGAUAGUGGAUCGUGGGGCUUCUG-GGCAGCAGG ----((((.-..)))).....(((.((((((.---((((.((((((.....)))------))).))))((((....(((((.....))))).))))....)-)))))))). ( -36.40) >DroEre_CAF1 91066 97 - 1 ----ACAGA-AACUGUUCCAACUGGCUGCCCC---GCCACUUUAUUACGCUGAU------AAGUUGGCGCCUAAUAACGAUAGUGGAUCGAGGGGUUUCUAGGGCAGAGGA ----.....-..(((((....((((..(((((---.((((((((((((((..(.------...)..)))..))))))....))))).....)))))..))))))))).... ( -31.00) >DroPer_CAF1 108216 108 - 1 UGCCACAAAUAACUGUUCCAACUGGUUGCCCCUCUGCCACUUUAUUACGCCAAUGGAAUCAAGUUGGCGCCUAAUAAUAAU---GCUUCGGGGCCGCAGGGGGGCCGAAGG ..((.............((....))..((((((((((....(((((((((((((........)))))))..))))))....---(((....))).))))))))))....)) ( -35.30) >consensus ____ACAGA_AACUGUUCCAACUGGCUGCCCC___GCCACUUUAUUACGCUGAU______AAGUUGGCGCCUAAUAACGAUAGUGGAUCGAGGGCCUUCUGGGGCAGAAGG ....((((....)))).....((..(((((((....((...(((((((((((((........)))))))..))))))((((.....)))).)).......))))))).)). (-19.21 = -19.72 + 0.50)

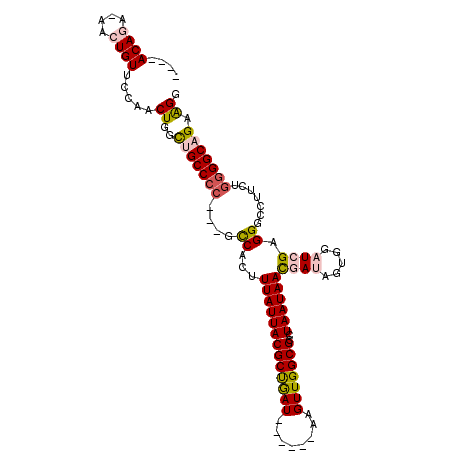
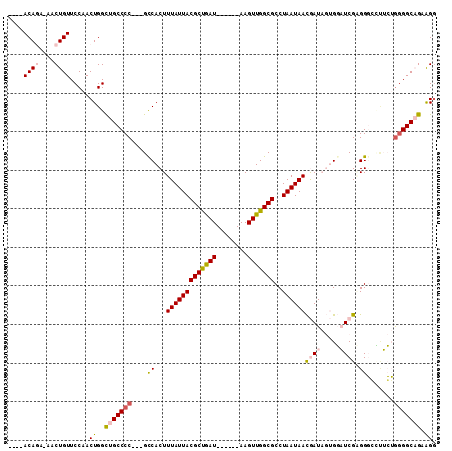
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:52:17 2006