
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,544,911 – 20,545,006 |
| Length | 95 |
| Max. P | 0.996821 |

| Location | 20,544,911 – 20,545,006 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 93.50 |
| Mean single sequence MFE | -28.80 |
| Consensus MFE | -24.73 |
| Energy contribution | -25.47 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.61 |
| SVM RNA-class probability | 0.967581 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20544911 95 + 23771897 AGCUUUGACUCGGUGGAUUUUUUUUACCCCUUUUGAUUAUCCCCACACCAAUGUGGUUGUAUCCGUGGAUUUCGGGGGAGUCAAAAGUCAAUUAA ....(((((..((((((.....))))))..((((((((.((((((((((.....)).)))(((....)))...)))))))))))))))))).... ( -28.80) >DroSec_CAF1 86832 94 + 1 AGCUUAGACUCAGUGGGUUU-UUUUACCCCCUUUGAUUAUCCCCACUCCAGUUUGGUUGUAUCCGUGGAUUUCGGGGGAGUCAAAAGUCAAUUAA ......((((..(.((((..-....)))).)(((((((.(((((..((((...(((......)))))))....))))))))))))))))...... ( -27.30) >DroSim_CAF1 87770 95 + 1 AGCUUUGACUCAGUGGGUUUUUUUUACCCCCUUUGAUUAUCCCCACUCCAGUUUGGUUGUAUCCGUGGAUUUCGGGGGAGUCAAAAGUCAAUUAA ....((((((..(.((((.......)))).)(((((((.(((((..((((...(((......)))))))....)))))))))))))))))).... ( -28.20) >DroEre_CAF1 89394 94 + 1 AGCUUUGACUCAGUGGGUUU-UUGUACCCUUUUUGAUUUUCCCCACUCCAAUGUGGUUGUAUCCGUGGAUUUCGGGGGAGUCAAAAGUCAAUUAA ....(((((.....((((..-....)))).((((((((..(((...((((...(((......)))))))....)))..))))))))))))).... ( -30.90) >consensus AGCUUUGACUCAGUGGGUUU_UUUUACCCCCUUUGAUUAUCCCCACUCCAAUGUGGUUGUAUCCGUGGAUUUCGGGGGAGUCAAAAGUCAAUUAA ....((((((....((((.......))))..(((((((.(((((..((((...(((......)))))))....)))))))))))))))))).... (-24.73 = -25.47 + 0.75)



| Location | 20,544,911 – 20,545,006 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 93.50 |
| Mean single sequence MFE | -27.70 |
| Consensus MFE | -26.36 |
| Energy contribution | -26.67 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.54 |
| Structure conservation index | 0.95 |
| SVM decision value | 2.75 |
| SVM RNA-class probability | 0.996821 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20544911 95 - 23771897 UUAAUUGACUUUUGACUCCCCCGAAAUCCACGGAUACAACCACAUUGGUGUGGGGAUAAUCAAAAGGGGUAAAAAAAAUCCACCGAGUCAAAGCU .........(((((((((((((...(((....)))(((.((.....)))))))))..........((((..........)).))))))))))).. ( -25.50) >DroSec_CAF1 86832 94 - 1 UUAAUUGACUUUUGACUCCCCCGAAAUCCACGGAUACAACCAAACUGGAGUGGGGAUAAUCAAAGGGGGUAAAA-AAACCCACUGAGUCUAAGCU ........(((..((((((((((...((((.((......))....)))).)))))........((.((((....-..)))).))))))).))).. ( -26.90) >DroSim_CAF1 87770 95 - 1 UUAAUUGACUUUUGACUCCCCCGAAAUCCACGGAUACAACCAAACUGGAGUGGGGAUAAUCAAAGGGGGUAAAAAAAACCCACUGAGUCAAAGCU .........((((((((((((((...((((.((......))....)))).)))))........((.((((.......)))).))))))))))).. ( -29.10) >DroEre_CAF1 89394 94 - 1 UUAAUUGACUUUUGACUCCCCCGAAAUCCACGGAUACAACCACAUUGGAGUGGGGAAAAUCAAAAAGGGUACAA-AAACCCACUGAGUCAAAGCU .........((((((((((((((...((((.((......))....)))).)))))...........((((....-..))))...))))))))).. ( -29.30) >consensus UUAAUUGACUUUUGACUCCCCCGAAAUCCACGGAUACAACCAAACUGGAGUGGGGAUAAUCAAAAGGGGUAAAA_AAACCCACUGAGUCAAAGCU .........((((((((((((((...((((.((......))....)))).)))))...........((((.......))))...))))))))).. (-26.36 = -26.67 + 0.31)

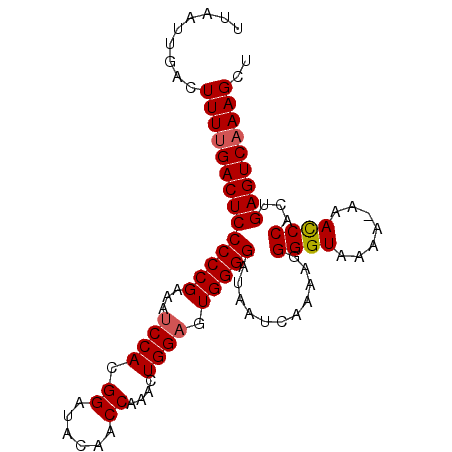

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:52:11 2006