
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,381,044 – 20,381,197 |
| Length | 153 |
| Max. P | 0.678156 |

| Location | 20,381,044 – 20,381,144 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 91.17 |
| Mean single sequence MFE | -33.92 |
| Consensus MFE | -25.71 |
| Energy contribution | -26.77 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.678156 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
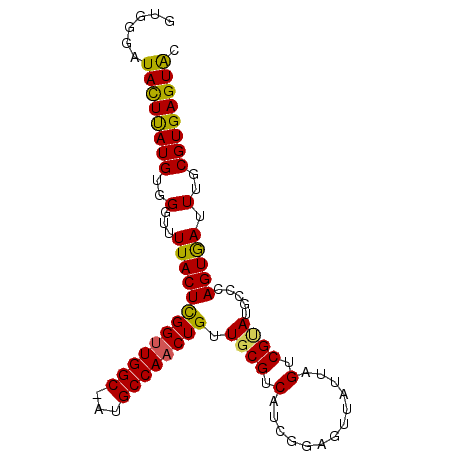
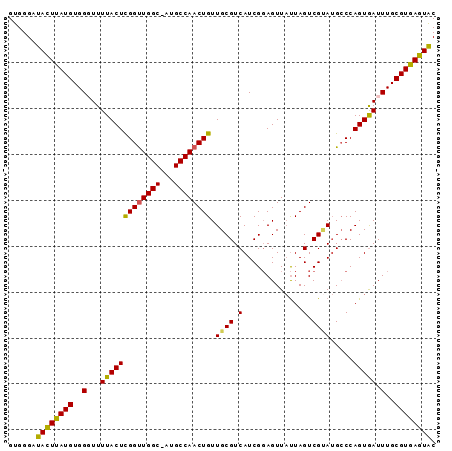
>3L_DroMel_CAF1 20381044 100 - 23771897 CGGAGUUAUUAGUCGAAUGGCCAGUGAUUUGCGUGAGUACGGCUUGUUAAGGGCCCAAAAUUAGCUAAUUGGGCUCUUUAGUGAGUUGCCGAUGCCACUG ..((((((((.(((....))).))))))))..(((.((((((((..((((((((((((..........)))))).))))))..)...)))).)))))).. ( -33.50) >DroSec_CAF1 9291 100 - 1 CGGAGUUAUUAGUCGUAUGCCCAGUGAUUUGCGUGAGUGCGGCUUGUUAAGGGCCCAAAAUUAGCUAAUUGGGCUCUUUAGUGAGUUGCCGCUGGCACUG .................((((.((((....((....))((((((..((((((((((((..........)))))).))))))..))))))))))))))... ( -37.70) >DroSim_CAF1 9473 100 - 1 CGGAGUUAUUAGUCGUAUGCCCAGUGAUUUGCGUGAGUGCGGCUUGUUAAGGGCCCAAAAUUAGCUAAUUGGGCUCUUUAGUGAGUUGCCGCUGCCACAG .((...(((.....)))..))(((((....((....))((((((..((((((((((((..........)))))).))))))..)))))))))))...... ( -33.50) >DroEre_CAF1 9165 99 - 1 CGCAGUUCCUAGUCGCAAGCCCAGUAAUUUGCGUGAGUACGGCUUGUUAAGGGC-CAAAAUUAGCUAAUUGGGCUCUUUAGAGAGUCGACUGUGCCACUG ..((((..((((((((..(((((((....(((....))).(((((.....))))-)...........)))))))(((....)))).)))))).)..)))) ( -31.00) >consensus CGGAGUUAUUAGUCGUAUGCCCAGUGAUUUGCGUGAGUACGGCUUGUUAAGGGCCCAAAAUUAGCUAAUUGGGCUCUUUAGUGAGUUGCCGCUGCCACUG .....................(((((....((....))((((((..((((((((((((..........)))))).))))))..))))))......))))) (-25.71 = -26.77 + 1.06)



| Location | 20,381,104 – 20,381,197 |
|---|---|
| Length | 93 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 87.46 |
| Mean single sequence MFE | -28.87 |
| Consensus MFE | -22.47 |
| Energy contribution | -21.97 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.22 |
| SVM RNA-class probability | 0.639788 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20381104 93 - 23771897 GUGGGAUACUUAUGUGGGUUUUACUCGGUUGGCUAUGCCAACUGUUGCGUCAUCGGAGUUAUUAGUCGAAUGGCCAGUGAUUUGCGUGAGUAC ......((((((((..((((.....((((((((...))))))))(((.(((((..((.(....).))..)))))))).))))..)))))))). ( -30.10) >DroSec_CAF1 9351 86 - 1 GUGGGAUACUUAUGUGGCUUUUACUCGGUUG-------CAACUGUUGCGUCAUCGGAGUUAUUAGUCGUAUGCCCAGUGAUUUGCGUGAGUGC ......((((((((..(...(((((.(((((-------(.((((..((.((....))))...)))).))).))).))))).)..)))))))). ( -23.70) >DroSim_CAF1 9533 93 - 1 GUGGGAUACUUAUGUGGGUUUUACUCGGUUGGCCAAGCCAACUGUUGCGUCAUCGGAGUUAUUAGUCGUAUGCCCAGUGAUUUGCGUGAGUGC ......((((((((..(........((((((((...))))))))....(((((.((.((............)))).))))))..)))))))). ( -26.40) >DroEre_CAF1 9224 93 - 1 GUGGGAUAUUCAUGUGGUUUUUACUUGGCUGGCGAUGCCAACUGUUGCGUCAUCGCAGUUCCUAGUCGCAAGCCCAGUAAUUUGCGUGAGUAC ......((((((((..(...(((((.((((.(((((...((((((.........))))))....))))).)))).))))).)..)))))))). ( -35.30) >consensus GUGGGAUACUUAUGUGGGUUUUACUCGGUUGGC_AUGCCAACUGUUGCGUCAUCGGAGUUAUUAGUCGUAUGCCCAGUGAUUUGCGUGAGUAC ......((((((((..(...(((((((((((((...)))))))).((((.(.............).)))).....))))).)..)))))))). (-22.47 = -21.97 + -0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:50:50 2006