| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,278,709 – 20,278,885 |
| Length | 176 |
| Max. P | 0.999314 |

| Location | 20,278,709 – 20,278,825 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.58 |
| Mean single sequence MFE | -24.02 |
| Consensus MFE | -18.50 |
| Energy contribution | -19.94 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.804269 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
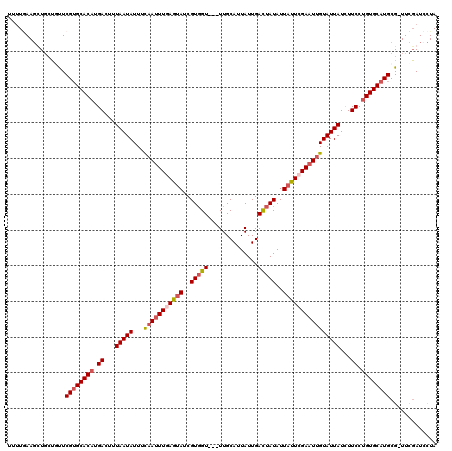
>3L_DroMel_CAF1 20278709 116 - 23771897 UUUUAAAGCUGAUUUUCGUGCACUUGACUUUAAUAUUUUAAUUUGAGUAUCGUGGU---UUGCAUUAUUGAUUAUAUUAUUCGAAUUGUAUUAUCUUCCUGUGCAUGCG-UUCGAUUCUA .........(((....(((((((..((...((((((...((((((((((..(((((---(.........))))))..))))))))))))))))...))..)))))))..-.)))...... ( -24.70) >DroSec_CAF1 17309 116 - 1 UUUUGAAGUUGCUUUUCGUGCACAUGACUUUAAUAUUUCAAUUUUAGUAUCGUGGU---UUGCAUUAUUGACUAUAUUACUCGAAUUGUAUUAUCUUCCUGUGCAUGCG-UUCGAUCCUA ..(((((..(((.....(((((((.((...(((((...((((((.((((..(((((---(.........))))))..)))).)))))))))))...)).))))))))))-)))))..... ( -26.50) >DroSim_CAF1 17273 116 - 1 UUUUGAAGCUGCUGUUCGUGCACAUGACUUUAAUAUUUCAAUUUUAGUAUCGUGGU---UUGCAUUAUUGACUAUAUUAUUCGAAUUGUAUUAUCUUCCUGUGCAUGCG-UUCGAUCCUA ..(((((((....)).((((((((.((...(((((...((((((.((((..(((((---(.........))))))..)))).)))))))))))...)).))))))))..-)))))..... ( -24.30) >DroYak_CAF1 8989 120 - 1 AAAUCAAUUUUCAGCUCGGGCACAUGACUUUAAUAUUUCAACUUGAGCAUCGUUGUUGCUUGCAUUAUUGACUAUAUUAUUCGAAUGGUAUUAUUUUCCUGUGCAUGCGGUUUUAUCUUA (((((........(((((((....(((..(....)..))).))))))).........((.(((((....((..(((((((....)))))))..)).....))))).)))))))....... ( -20.60) >consensus UUUUGAAGCUGCUGUUCGUGCACAUGACUUUAAUAUUUCAAUUUGAGUAUCGUGGU___UUGCAUUAUUGACUAUAUUAUUCGAAUUGUAUUAUCUUCCUGUGCAUGCG_UUCGAUCCUA ................((((((((.((...(((((...(((((((((((..(((((..............)))))..))))))))))))))))...)).))))))))............. (-18.50 = -19.94 + 1.44)


| Location | 20,278,785 – 20,278,885 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.60 |
| Mean single sequence MFE | -28.50 |
| Consensus MFE | -19.52 |
| Energy contribution | -20.40 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.45 |
| Structure conservation index | 0.69 |
| SVM decision value | 3.51 |
| SVM RNA-class probability | 0.999314 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
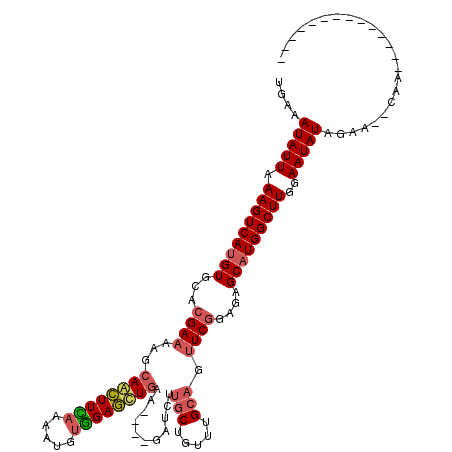
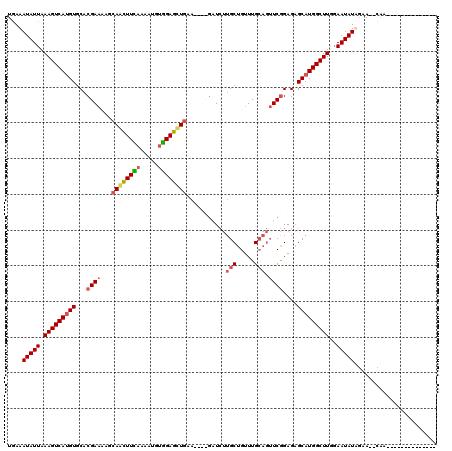
>3L_DroMel_CAF1 20278785 100 + 23771897 UAAAAUAUUAAAGUCAAGUGCACGAAAAUCAGCUUUAAAAUGUGGAGCUGUA----GGUCUUGCUGUUUGCAGUUCGGAGAGCAUGGCUUGGAAUAUUGAA--CAA-------------- ...((((((.((((((.((...((((...((((((((.....))))))))..----....((((.....))))))))....)).))))))..))))))...--...-------------- ( -25.40) >DroSec_CAF1 17385 104 + 1 UGAAAUAUUAAAGUCAUGUGCACGAAAAGCAACUUCAAAAUGUGGAGCUGAAAGUUGAUCUUGCUGUUUGCAGUUCGGAGAGCAUGGCUUGGAAUAUUGAG--CAA-------------- ((.((((((.(((((((((...((((..(((((((((.....))))).....(((.......)))..))))..))))....)))))))))..))))))...--)).-------------- ( -27.70) >DroSim_CAF1 17349 104 + 1 UGAAAUAUUAAAGUCAUGUGCACGAACAGCAGCUUCAAAAUGUGGAGCUGAAAGUUGAUCUUGCUGUUUGCAGUUCGGAGAGCAUGGCUUGGAAUAUAGAA--CAA-------------- ....(((((.(((((((((...(((((..((((((((.....))))))))...........(((.....))))))))....)))))))))..)))))....--...-------------- ( -34.80) >DroYak_CAF1 9069 111 + 1 UGAAAUAUUAAAGUCAUGUGCCCGAGCUGAAAAUUGAUUUUGCUGAUUUGU------GUGGUGCUGUUUG---GUCAGAGCGCAUGGCUUUCAAUAUAUAGAGUUAAAGUUUAUAAUAGU ....(((((((((((((((((.(..((..((....(...(..(........------)..)..)..))..---))..).)))))))))))).)))))....................... ( -26.10) >consensus UGAAAUAUUAAAGUCAUGUGCACGAAAAGCAACUUCAAAAUGUGGAGCUGAA____GAUCUUGCUGUUUGCAGUUCGGAGAGCAUGGCUUGGAAUAUAGAA__CAA______________ ....(((((.(((((((((...((((...((((((((.....))))))))...........(((.....))).))))....)))))))))..)))))....................... (-19.52 = -20.40 + 0.88)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:50:04 2006