
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,253,353 – 20,253,671 |
| Length | 318 |
| Max. P | 0.956909 |

| Location | 20,253,353 – 20,253,471 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.32 |
| Mean single sequence MFE | -29.15 |
| Consensus MFE | -25.43 |
| Energy contribution | -26.06 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.891481 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20253353 118 + 23771897 AUGCCCAAGUUUGUGACUGCCACAGAUUUCACUCUUAAGGGAGUCCUCAUCCAGGUGACAGAACUUCUAAAUCCAAUCUGGAAAU--ACAUUUGUAAAUAACUUCUAAUUCUAGAUGGGC ..(((((((((((((.....))))))))..........((((((.....(((((((...................)))))))..(--((....)))....)))))).........))))) ( -27.11) >DroSec_CAF1 5658 118 + 1 AUGCCCAAGUUUGUGACUGCCACAGAUUUCACUCUUAAGGGAGUCCUCAUCCAGGUGCGAGAACUUCUAAAUCCAAUCUGGAAAU--ACAUUUGUAAAUAACUUCUAAUUCUAGAUGGGC ..(((((((((((((.....))))))))..........((((((.....(((((((...((.....)).......)))))))..(--((....)))....)))))).........))))) ( -27.80) >DroSim_CAF1 5634 118 + 1 AUGCCCAAGUUUGUGACUGCCACAGAUUUCACUCUUAAGGGAGUCCUCAUCCAGGUGCGAGAACUUCUAAAUCCAAUCUGGAAAU--ACAUUUGUAAAUAACUUCUAAUUCUAGAUGGGC ..(((((((((((((.....))))))))..........((((((.....(((((((...((.....)).......)))))))..(--((....)))....)))))).........))))) ( -27.80) >DroYak_CAF1 6176 119 + 1 AUGCCCAAGUUUGUCACUUCCACAGAUUUCACUCUUAAGGAAGUCCUUAUCGAGGUGGGAGAACUUCUAAAUCUAAUCUGGACCUUAAUAUUUG-AAAUACAUUUCGAUUCCAGAUGGGC ..((((..........(((((((.(....).(((.(((((....)))))..))))))))))..............(((((((.........(((-(((....)))))).))))))))))) ( -33.90) >consensus AUGCCCAAGUUUGUGACUGCCACAGAUUUCACUCUUAAGGGAGUCCUCAUCCAGGUGCGAGAACUUCUAAAUCCAAUCUGGAAAU__ACAUUUGUAAAUAACUUCUAAUUCUAGAUGGGC ..(((((((((((((.....))))))))).........((((((((((((....))))).).)))))).......((((((((.........................)))))))))))) (-25.43 = -26.06 + 0.63)


| Location | 20,253,471 – 20,253,591 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.72 |
| Mean single sequence MFE | -43.70 |
| Consensus MFE | -39.62 |
| Energy contribution | -40.06 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.758981 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
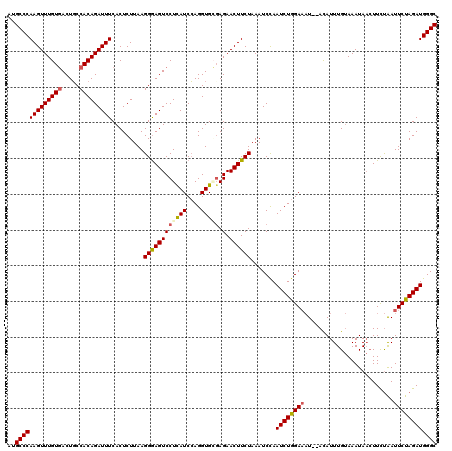
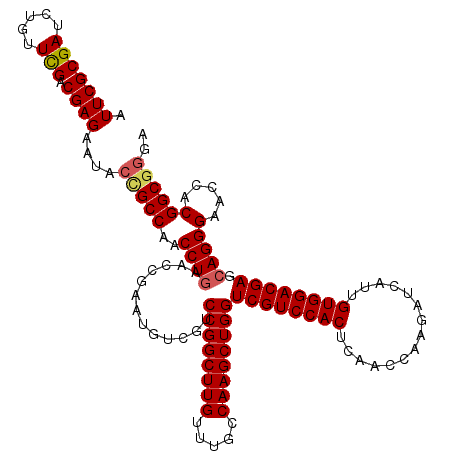
>3L_DroMel_CAF1 20253471 120 + 23771897 AUUCGCGAUCUGUUCGACGAGAAUACAGCCAACCUGGACCGAAUGUCUUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAACAGGGAACCACGGCGGGA .(((((((.....))).)))).......((..(((((.((.........((((((((.....))))))))((((((((...............))))))))...))...))).))..)). ( -41.16) >DroSec_CAF1 5776 120 + 1 AUUCGCGAUCUGUUCGACGAGAAUACCGCCAACCUGAACCGAAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGA .(((((((.....))).))))....(((((.........((.....))((((((......)))...(((.((((((((...............)))))))).)))))).....))))).. ( -44.66) >DroSim_CAF1 5752 120 + 1 AUUCGCGAUCUGUUCGACGAGAAUACCGCCAACCUGAACCGAAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGA .(((((((.....))).))))....(((((.........((.....))((((((......)))...(((.((((((((...............)))))))).)))))).....))))).. ( -44.66) >DroYak_CAF1 6295 120 + 1 AUUCGCGAUCUGUUUGACGAGAACAAUGCCAACCUAAGCCGGAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCGACCAAGAUCAUCGUGGACGAGCAGGGAACCACGGCGGGA .(((((((((((((((.((((.............((((((((((...)))))))))))))).)))))((((((......)))))))))))..(((((.(......)...)))))))))). ( -44.31) >consensus AUUCGCGAUCUGUUCGACGAGAAUACCGCCAACCUGAACCGAAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGA .(((((((.....))).))))....(((((..((((.............((((((((.....))))))))((((((((...............)))))))).))))(.....)))))).. (-39.62 = -40.06 + 0.44)


| Location | 20,253,511 – 20,253,631 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.08 |
| Mean single sequence MFE | -50.64 |
| Consensus MFE | -49.47 |
| Energy contribution | -49.53 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.98 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.815556 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
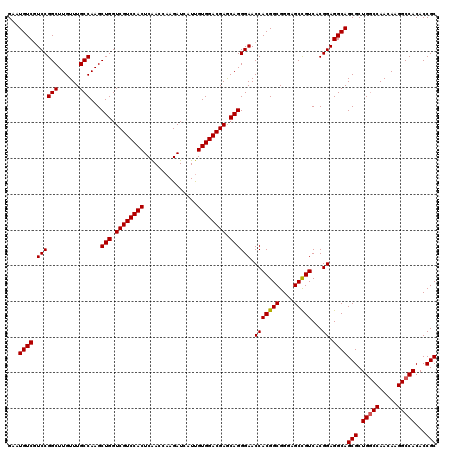
>3L_DroMel_CAF1 20253511 120 + 23771897 GAAUGUCUUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAACAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGC ...(((((((((((......)))...(((.((((((((...............)))))))).)))))).(((((((....)))))...)))))))(((.(((((.....)))))...))) ( -50.66) >DroSec_CAF1 5816 120 + 1 GAAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGC ...((((.((((((......)))...(((.((((((((...............)))))))).)))))).(((((((....)))))...)).))))(((.(((((.....)))))...))) ( -52.56) >DroSim_CAF1 5792 120 + 1 GAAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGC ...((((.((((((......)))...(((.((((((((...............)))))))).)))))).(((((((....)))))...)).))))(((.(((((.....)))))...))) ( -52.56) >DroYak_CAF1 6335 120 + 1 GGAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCGACCAAGAUCAUCGUGGACGAGCAGGGAACCACGGCGGGAGCUGUCACGGAGGCAGCGCUGUCCAACAAGGCCACACCGC ((((...))))((((((((.(((...(((.((((((((..((......))...)))))))).)))(....)..)))(((.((((((.....))))))....)))))))).)))....... ( -46.80) >consensus GAAUGUCGUCCGGCUUGUUUGCCAAGCUGGUCGUCCACUCAACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGC ...((((.((((((......)))...(((.((((((((...............)))))))).)))))).(((((((....)))))...)).))))(((.(((((.....)))))...))) (-49.47 = -49.53 + 0.06)


| Location | 20,253,551 – 20,253,671 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 96.53 |
| Mean single sequence MFE | -46.18 |
| Consensus MFE | -43.30 |
| Energy contribution | -44.05 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.830825 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
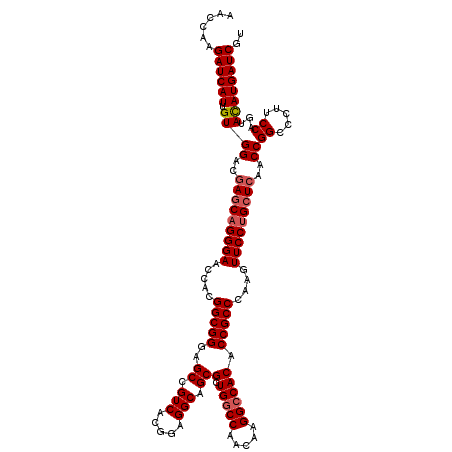
>3L_DroMel_CAF1 20253551 120 + 23771897 AACCAAGAUCAUUGUGGACGAACAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGCCCAAGUUCCUGCUCAACCGGCCCUUCCAGUACAUGAUCGU ......((((((.((((..((.((((((.....(((((..(((........)))...(.(((((.....)))))).)))))....)))))).))..))((.....))...)))))))).. ( -44.20) >DroSec_CAF1 5856 120 + 1 AACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGCCCAAGUUCCUGCUCAACCGGCCCUUCCAGUAUAUGAUCGU ......((((((...((..(((((((((.....(((((..(((........)))...(.(((((.....)))))).)))))....)))))))))..))((.....)).....)))))).. ( -49.70) >DroSim_CAF1 5832 120 + 1 AACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGCCCAAGUUCCUGCUCAACCGGCCCUUCCAGUAUAUGAUCGU ......((((((...((..(((((((((.....(((((..(((........)))...(.(((((.....)))))).)))))....)))))))))..))((.....)).....)))))).. ( -49.70) >DroYak_CAF1 6375 120 + 1 GACCAAGAUCAUCGUGGACGAGCAGGGAACCACGGCGGGAGCUGUCACGGAGGCAGCGCUGUCCAACAAGGCCACACCGCCCAAGUUCCAGCUGAACCGGCCCUUCCAGUACAUGAUCGU ......((((((..((((.(.....(((((...(((((..((((((.....))))))(.((.((.....)).))).)))))...))))).((((...)))).).))))....)))))).. ( -41.10) >consensus AACCAAGAUCAUUGUGGACGAGCAGGGAACCACGGCGGGAGCCGUCACGGAGGCAGCGCUGGCCAACAAGGCCACACCGCCCAAGUUCCUGCUCAACCGGCCCUUCCAGUACAUGAUCGU ......((((((.((((..(((((((((.....(((((..((.(((.....))).))(.(((((.....)))))).)))))....)))))))))..))((.....))...)))))))).. (-43.30 = -44.05 + 0.75)


| Location | 20,253,551 – 20,253,671 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.53 |
| Mean single sequence MFE | -55.10 |
| Consensus MFE | -51.31 |
| Energy contribution | -52.12 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.02 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.956909 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
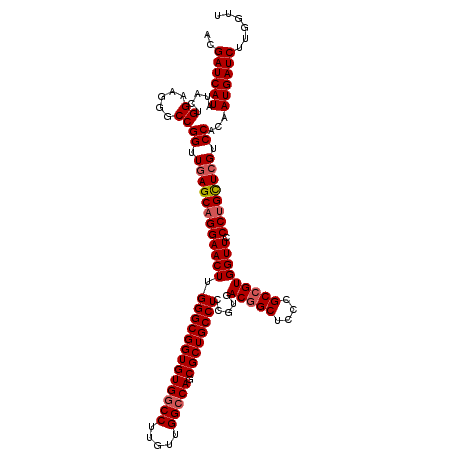
>3L_DroMel_CAF1 20253551 120 - 23771897 ACGAUCAUGUACUGGAAGGGCCGGUUGAGCAGGAACUUGGGCGGUGUGGCCUUGUUGGCCAGCGCUGCCUCCGUGACGGCUCCCGCCGUGGUUCCCUGUUCGUCCACAAUGAUCUUGGUU ..((((((((...((.....))((.((((((((((((.(((((((((((((.....))))).)))))))).....(((((....))))))))).)))))))).)))).))))))...... ( -55.90) >DroSec_CAF1 5856 120 - 1 ACGAUCAUAUACUGGAAGGGCCGGUUGAGCAGGAACUUGGGCGGUGUGGCCUUGUUGGCCAGCGCUGCCUCCGUGACGGCUCCCGCCGUGGUUCCCUGCUCGUCCACAAUGAUCUUGGUU ..((((((.....((.....))((.((((((((((((.(((((((((((((.....))))).)))))))).....(((((....))))))))).)))))))).))...))))))...... ( -57.80) >DroSim_CAF1 5832 120 - 1 ACGAUCAUAUACUGGAAGGGCCGGUUGAGCAGGAACUUGGGCGGUGUGGCCUUGUUGGCCAGCGCUGCCUCCGUGACGGCUCCCGCCGUGGUUCCCUGCUCGUCCACAAUGAUCUUGGUU ..((((((.....((.....))((.((((((((((((.(((((((((((((.....))))).)))))))).....(((((....))))))))).)))))))).))...))))))...... ( -57.80) >DroYak_CAF1 6375 120 - 1 ACGAUCAUGUACUGGAAGGGCCGGUUCAGCUGGAACUUGGGCGGUGUGGCCUUGUUGGACAGCGCUGCCUCCGUGACAGCUCCCGCCGUGGUUCCCUGCUCGUCCACGAUGAUCUUGGUC ..((((((....((((.((((((((..(((((..((..(((((((((..((.....))...)))))))))..))..)))))...)))).((...)).)))).))))..))))))...... ( -48.90) >consensus ACGAUCAUAUACUGGAAGGGCCGGUUGAGCAGGAACUUGGGCGGUGUGGCCUUGUUGGCCAGCGCUGCCUCCGUGACGGCUCCCGCCGUGGUUCCCUGCUCGUCCACAAUGAUCUUGGUU ..((((((.....((.....))((.((((((((((((.(((((((((((((.....))))).)))))))).....(((((....))))))))).)))))))).))...))))))...... (-51.31 = -52.12 + 0.81)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:49:58 2006