
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,236,708 – 20,236,840 |
| Length | 132 |
| Max. P | 0.969127 |

| Location | 20,236,708 – 20,236,800 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 97.50 |
| Mean single sequence MFE | -18.72 |
| Consensus MFE | -17.38 |
| Energy contribution | -17.66 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.770457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20236708 92 + 23771897 UCUGGUAGACACACAUAUAUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUG ...((.(..((((...((((((...(((((((((..........)))))))))....))))))..))))..).))(((((.....))))).. ( -18.90) >DroSec_CAF1 6269 92 + 1 UCUGGUAGACACACAUAUAUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUG ...((.(..((((...((((((...(((((((((..........)))))))))....))))))..))))..).))(((((.....))))).. ( -18.90) >DroSim_CAF1 7513 92 + 1 UCUGGUAGACACACAUAUAUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUG ...((.(..((((...((((((...(((((((((..........)))))))))....))))))..))))..).))(((((.....))))).. ( -18.90) >DroEre_CAF1 7058 89 + 1 UCUGGUAGACA--CAUAUAUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUUCCGUGAAACCU-UUCGCUG ...((....((--((.((((((...(((((((((..........)))))))))....))))))..))))....))(((((....-))))).. ( -18.80) >DroYak_CAF1 11577 91 + 1 UCUGGUAGACACGCAUAUAUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUUCCGUGAAACCU-UUCGCUG ...((....((((...((((((...(((((((((..........)))))))))....))))))..))))....))(((((....-))))).. ( -18.10) >consensus UCUGGUAGACACACAUAUAUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUG ...((.(..((((...((((((...(((((((((..........)))))))))....))))))..))))..).))(((((.....))))).. (-17.38 = -17.66 + 0.28)



| Location | 20,236,708 – 20,236,800 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 97.50 |
| Mean single sequence MFE | -25.76 |
| Consensus MFE | -22.26 |
| Energy contribution | -23.26 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938337 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20236708 92 - 23771897 CAGCGAAGAGGUUUCACGGGAUGCACAGGACAUCGCGGUAUACAUAUAGUGGCUAUAGUAUAUAUAUGGUCGAUAUAUGUGUGUCUACCAGA .........(((.....(.(((((....).)))).)(..(((((((((.((((((((.......)))))))).)))))))))..).)))... ( -26.90) >DroSec_CAF1 6269 92 - 1 CAGCGAAGAGGUUUCACGGGAUGCACAGGACAUCGCGGUAUACAUAUAGUGGCUAUAGUAUAUAUAUGGUCGAUAUAUGUGUGUCUACCAGA .........(((.....(.(((((....).)))).)(..(((((((((.((((((((.......)))))))).)))))))))..).)))... ( -26.90) >DroSim_CAF1 7513 92 - 1 CAGCGAAGAGGUUUCACGGGAUGCACAGGACAUCGCGGUAUACAUAUAGUGGCUAUAGUAUAUAUAUGGUCGAUAUAUGUGUGUCUACCAGA .........(((.....(.(((((....).)))).)(..(((((((((.((((((((.......)))))))).)))))))))..).)))... ( -26.90) >DroEre_CAF1 7058 89 - 1 CAGCGAA-AGGUUUCACGGAAUGCACAGGACAUCGCGGUAUACAUAUAGUGGCUAUAGUAUAUAUAUGGUCGAUAUAUG--UGUCUACCAGA ..((((.-..((((.............)))).))))((((.(((((((.((((((((.......))))))))...))))--))).))))... ( -25.62) >DroYak_CAF1 11577 91 - 1 CAGCGAA-AGGUUUCACGGAAUGCACAGGACAUCGCGGUAUACAUAUAGUGGCUAUAGUAUAUAUAUGGUCGAUAUAUGCGUGUCUACCAGA .......-.(((..((((..((((...(.....)...)))).((((((.((((((((.......)))))))).))))))))))...)))... ( -22.50) >consensus CAGCGAAGAGGUUUCACGGGAUGCACAGGACAUCGCGGUAUACAUAUAGUGGCUAUAGUAUAUAUAUGGUCGAUAUAUGUGUGUCUACCAGA .........(((.....(.(((((....).)))).)(..(((((((((.((((((((.......)))))))).)))))))))..).)))... (-22.26 = -23.26 + 1.00)


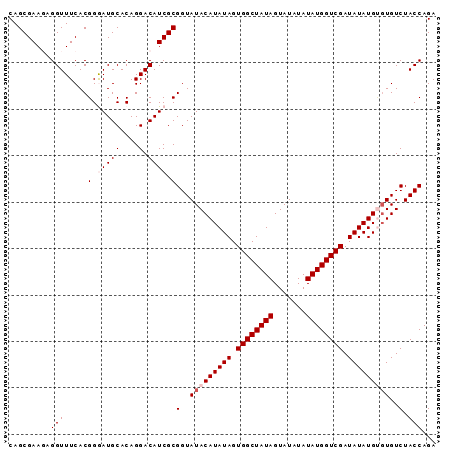
| Location | 20,236,726 – 20,236,840 |
|---|---|
| Length | 114 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 98.60 |
| Mean single sequence MFE | -33.60 |
| Consensus MFE | -33.64 |
| Energy contribution | -33.48 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.58 |
| Structure conservation index | 1.00 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969127 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20236726 114 + 23771897 AUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUGCAUCCGCGUAUCGGGGGUGGUCGGUCUGCCCGAAAAACUC (((((((((...((((((((((...))))).)))))((.(((((..(.(((((....(((((.....))))))))))..)..))))).)))))))))))............... ( -34.10) >DroSec_CAF1 6287 114 + 1 AUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUGCAUCCGCGUAUCGGGGGUGGUCGGUCUGCCCGAAAAACUC (((((((((...((((((((((...))))).)))))((.(((((..(.(((((....(((((.....))))))))))..)..))))).)))))))))))............... ( -34.10) >DroSim_CAF1 7531 114 + 1 AUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUGCAUCCGCGUAUCGGGGGUGGUCGGUCUGCCCGAAAAACUC (((((((((...((((((((((...))))).)))))((.(((((..(.(((((....(((((.....))))))))))..)..))))).)))))))))))............... ( -34.10) >DroEre_CAF1 7074 113 + 1 AUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUUCCGUGAAACCU-UUCGCUGCAUCCGCGUAUUGGGGGUGGUCGGUCUGCCCGAAAAACUC (((((((((...((((((((((...))))).)))))((.(((((..(.(((((....(((((....-))))))))))..)..))))).)))))))))))............... ( -31.80) >DroYak_CAF1 11595 113 + 1 AUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUUCCGUGAAACCU-UUCGCUGCAUCCGCGUAUCGGGGGUGGUCGGUCUGCCCGAAAAACUC (((((((((...((((((((((...))))).)))))((.(((((..(.(((((....(((((....-))))))))))..)..))))).)))))))))))............... ( -33.90) >consensus AUCGACCAUAUAUAUACUAUAGCCACUAUAUGUAUACCGCGAUGUCCUGUGCAUCCCGUGAAACCUCUUCGCUGCAUCCGCGUAUCGGGGGUGGUCGGUCUGCCCGAAAAACUC (((((((((...((((((((((...))))).)))))((.(((((..(.(((((....(((((.....))))))))))..)..))))).)))))))))))............... (-33.64 = -33.48 + -0.16)

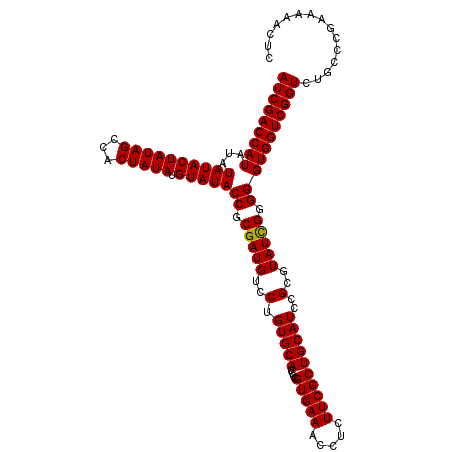

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:49:39 2006