
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,172,785 – 20,172,908 |
| Length | 123 |
| Max. P | 0.864053 |

| Location | 20,172,785 – 20,172,884 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 77.46 |
| Mean single sequence MFE | -30.89 |
| Consensus MFE | -18.75 |
| Energy contribution | -18.45 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.61 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.864053 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
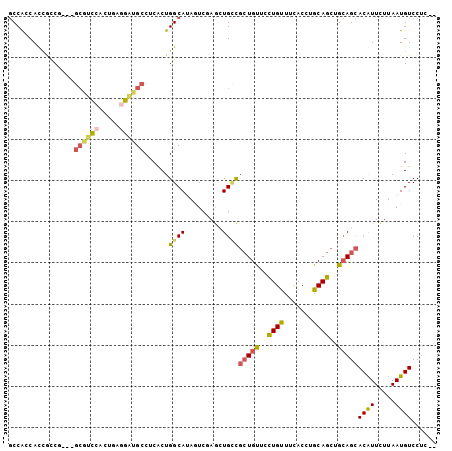
>3L_DroMel_CAF1 20172785 99 + 23771897 GCCACCACCACCG---GCGUCCACUGAGGAUGCCUCACUGGCAAAAUCGAGCUGCCGAUGUUCCUGUUUCACCUGCAGCUGCAGCACGUUCUUAAUGUCCUC-- ((((........(---((((((.....)))))))....))))......(((((((...(((..((((.......))))..)))))).))))...........-- ( -28.50) >DroPse_CAF1 5368 102 + 1 GCCGGCAUCGCCUCCCGCUGUUGCCGCCGCAGCCUCACUGGCAUAGUCUAAGUGCUGCUGCUCCUGUUUCACUUGCAGCUGCAGCACAUUUUUAAUGUCCUC-- (((((((..((.....))...)))))..)).(((.....)))(((....(((((.((((((..((((.......))))..)))))))))))....)))....-- ( -33.10) >DroSim_CAF1 6031 99 + 1 GCCACCACCACCG---GCGUCCACUGAGGAUGCCUCACUGGCAAAAUCGAGCUGCCGCUGUUCUUGUUUCACCUGCAGAUGCAGCACGUUCUUAAUGUCCUC-- ((((........(---((((((.....)))))))....)))).....((.(((((..((((.............))))..))))).))..............-- ( -29.62) >DroYak_CAF1 5071 99 + 1 GCCACCACCGCCA---GCGUCCACUGAGGAUGCCUCACUGGCAUAUUCAAGCUGUCGCUGUUCCUGUUUUACUUGCAGCUGCAGCACGUUCUUAAUGUCCUC-- .........((((---((((((.....)))))).....))))........(((((.(((((.............))))).))))).................-- ( -29.12) >DroAna_CAF1 5819 98 + 1 UCCGCCACUCUC------CACCGCAGCGGAUGGCUCGCUGGCAUAGUCCAGCUGGCGCUGUUCCUGCUUCACCUGCAGCUGAAGGACAUUCUUAAUGUCCUCUC (((((..(....------....)..)))))..(((.(((((......))))).)))(((((.............)))))...(((((((.....)))))))... ( -31.92) >DroPer_CAF1 5361 102 + 1 GCCGGCAUCGCCUCCCGCUGUUGCCGCCGCAGCCUCACUGGCAUAGUCUAAGUGCUGCUGCUCCUGUUUCACUUGCAGCUGCAGCACAUUUUUAAUGUCCUC-- (((((((..((.....))...)))))..)).(((.....)))(((....(((((.((((((..((((.......))))..)))))))))))....)))....-- ( -33.10) >consensus GCCACCACCGCCG___GCGUCCACUGAGGAUGCCUCACUGGCAUAGUCGAGCUGCCGCUGUUCCUGUUUCACCUGCAGCUGCAGCACAUUCUUAAUGUCCUC__ ................((((((.....))))))......((((.........))))(((((..((((.......))))..)))))((((.....))))...... (-18.75 = -18.45 + -0.30)


| Location | 20,172,809 – 20,172,908 |
|---|---|
| Length | 99 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 78.11 |
| Mean single sequence MFE | -24.67 |
| Consensus MFE | -15.47 |
| Energy contribution | -15.22 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.659201 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20172809 99 + 23771897 GGAUGCCUCACUGGCAAAAUCGAGCUGCCGAUGUUCCUGUUUCACCUGCAGCUGCAGCACGUUCUUAAUGUCCUCCUGCCCCGCUCCAUUGUCCUCCAG (((((((.....))).....((((((((.(.((.........)).).))))))((((.((((.....))))....))))..)).......))))..... ( -22.10) >DroVir_CAF1 10872 90 + 1 AGCUGCAUCAUUGGCAUAGUCCAGCUGCCGAUGCUCUUGCUUGACGUGCAACUGCAACACACUUUUAAUCUCCUCCUGCCC---------AUCCUCUGG .(.((((.((((((((.........))))))))...((((.......)))).)))).).....................((---------(.....))) ( -16.10) >DroPse_CAF1 5395 99 + 1 CGCAGCCUCACUGGCAUAGUCUAAGUGCUGCUGCUCCUGUUUCACUUGCAGCUGCAGCACAUUUUUAAUGUCCUCCUGGCCGCCUGUUCCCUCUUCCAA .((.(((.....(((((.....(((((.((((((..((((.......))))..)))))))))))...))))).....))).))................ ( -30.00) >DroSim_CAF1 6055 99 + 1 GGAUGCCUCACUGGCAAAAUCGAGCUGCCGCUGUUCUUGUUUCACCUGCAGAUGCAGCACGUUCUUAAUGUCCUCCUGCCCCGCUCCAUCGUCAUCCAG (((((((.....)))......((((.((.(((((..((((.......))))..)))))((((.....))))......))...)))).......)))).. ( -25.10) >DroYak_CAF1 5095 99 + 1 GGAUGCCUCACUGGCAUAUUCAAGCUGUCGCUGUUCCUGUUUUACUUGCAGCUGCAGCACGUUCUUAAUGUCCUCCUGCCCCGCUCCAUCGUCCUCCAG (((((.......((((.......(((((.(((((.............))))).)))))((((.....)))).....)))).........)))))..... ( -24.71) >DroPer_CAF1 5388 99 + 1 CGCAGCCUCACUGGCAUAGUCUAAGUGCUGCUGCUCCUGUUUCACUUGCAGCUGCAGCACAUUUUUAAUGUCCUCCUGGCCGCCUGUUCCCUCUUCCAA .((.(((.....(((((.....(((((.((((((..((((.......))))..)))))))))))...))))).....))).))................ ( -30.00) >consensus GGAUGCCUCACUGGCAUAGUCGAGCUGCCGCUGCUCCUGUUUCACUUGCAGCUGCAGCACAUUCUUAAUGUCCUCCUGCCCCGCUCCAUCGUCCUCCAG ....(((.....)))..............(((((..((((.......))))..)))))......................................... (-15.47 = -15.22 + -0.25)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:49:01 2006