
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,093,147 – 20,093,270 |
| Length | 123 |
| Max. P | 0.978015 |

| Location | 20,093,147 – 20,093,241 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 92.70 |
| Mean single sequence MFE | -34.36 |
| Consensus MFE | -27.60 |
| Energy contribution | -27.88 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.80 |
| SVM RNA-class probability | 0.978015 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20093147 94 + 23771897 AGGGCGUUGCAGGACAAUGAAAAGUGUUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAA------AACUGGGCAAC ...((((((((((.((((......(((.(((((....)))))))).......))))))))))))..(((.(((.....)))..------.)))..))... ( -33.52) >DroSec_CAF1 7736 100 + 1 AGGGCGUUGCAGGACAAUGAAAAGUGUUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAAUGUGGCAACUAGGCAAC ...((((((((((.((((......(((.(((((....)))))))).......))))))))))))..(((.((((.(........))))).)))..))... ( -36.32) >DroSim_CAF1 7990 100 + 1 AGGGCGUUGCAGGACAAUGAAAAGUGUUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAAUGUGGCAACUAGGCAAC ...((((((((((.((((......(((.(((((....)))))))).......))))))))))))..(((.((((.(........))))).)))..))... ( -36.32) >DroEre_CAF1 7639 99 + 1 CGCGCGUUGCAGGACAAUGAAAAGUGUUUGGGCAUUCGCCGGGCAAAAAUAAAUUGCUUGCAGCGG-GCGGCCAACUGGGUAAUGUGGCAACUAGGCAAC (((.(((((((((.((((......(((((((.......))))))).......))))))))))))).-)))((((.(........)))))........... ( -36.92) >DroYak_CAF1 7762 100 + 1 CGGGCGUUGCAGGACAAUGAAAAGUGUUCAGACAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUUGGUAAUGUGGCAACUAGGCAAC .....(((((.(((((.(....).)))))......(((((((((((.......)))))..))))))(((.((((.(........))))).)))..))))) ( -28.70) >consensus AGGGCGUUGCAGGACAAUGAAAAGUGUUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAAUGUGGCAACUAGGCAAC ....(((((((((.((((......(((.(((((....)))))))).......))))))))))))).....(((.....(((.........))).)))... (-27.60 = -27.88 + 0.28)

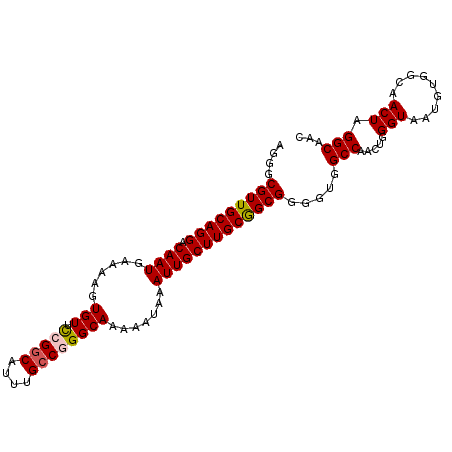

| Location | 20,093,147 – 20,093,241 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 92.70 |
| Mean single sequence MFE | -24.12 |
| Consensus MFE | -20.26 |
| Energy contribution | -20.54 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.585056 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20093147 94 - 23771897 GUUGCCCAGUU------UUACCCAGUUGGCCACCCCGCCGCAAGCAAUUUAUUUUUGCCCGGCAAAUGCCGGAACACUUUUCAUUGUCCUGCAACGCCCU (((((......------..........(((......)))((((((((.......))))(((((....)))))...........))))...)))))..... ( -24.40) >DroSec_CAF1 7736 100 - 1 GUUGCCUAGUUGCCACAUUACCCAGUUGGCCACCCCGCCGCAAGCAAUUUAUUUUUGCCCGGCAAAUGCCGGAACACUUUUCAUUGUCCUGCAACGCCCU (((((...((.(((((........).)))).))......((((((((.......))))(((((....)))))...........))))...)))))..... ( -25.70) >DroSim_CAF1 7990 100 - 1 GUUGCCUAGUUGCCACAUUACCCAGUUGGCCACCCCGCCGCAAGCAAUUUAUUUUUGCCCGGCAAAUGCCGGAACACUUUUCAUUGUCCUGCAACGCCCU (((((...((.(((((........).)))).))......((((((((.......))))(((((....)))))...........))))...)))))..... ( -25.70) >DroEre_CAF1 7639 99 - 1 GUUGCCUAGUUGCCACAUUACCCAGUUGGCCGC-CCGCUGCAAGCAAUUUAUUUUUGCCCGGCGAAUGCCCAAACACUUUUCAUUGUCCUGCAACGCGCG ...((...(((((.(((......((((((.((.-.(((((...((((.......)))).)))))..)).)))...)))......)))...)))))..)). ( -23.10) >DroYak_CAF1 7762 100 - 1 GUUGCCUAGUUGCCACAUUACCAAGUUGGCCACCCCGCCGCAAGCAAUUUAUUUUUGCCCGGCAAAUGUCUGAACACUUUUCAUUGUCCUGCAACGCCCG (((((...((.(((((........).)))).))...((((...((((.......)))).))))...........................)))))..... ( -21.70) >consensus GUUGCCUAGUUGCCACAUUACCCAGUUGGCCACCCCGCCGCAAGCAAUUUAUUUUUGCCCGGCAAAUGCCGGAACACUUUUCAUUGUCCUGCAACGCCCU (((((......................(((......)))((((((((.......))))(((((....)))))...........))))...)))))..... (-20.26 = -20.54 + 0.28)



| Location | 20,093,173 – 20,093,270 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 78.82 |
| Mean single sequence MFE | -37.70 |
| Consensus MFE | -20.84 |
| Energy contribution | -21.60 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.609276 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20093173 97 + 23771897 UUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAA------AACUGGGCAACUUGGC---UAG--------GCAACCAGGCAACCAGGCAUC .((((((....))))))...........((((((..((..(((.(((..(..(....------..)..)((......))---..)--------)).)))..))...)))))).. ( -35.10) >DroSec_CAF1 7762 114 + 1 UUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAAUGUGGCAACUAGGCAACUGGGCAACUAGGAAACCCGGCAACCCGUCAGCCAGGCAUC .((((((....))))))...........(((((((((((((...(((....((((..(.(((...((((....))))....))).)..)))))))..)))))).).)))))).. ( -47.50) >DroSim_CAF1 8016 114 + 1 UUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAAUGUGGCAACUAGGCAACUGGGCAACUAGGCAACUAGGCAACCCGGCAGCCGGGCAUC .((((((...(((((((..........((((((((.(((...((.((....)).)).))).))))((((....))))))))((((....))))....))))))))))))).... ( -51.70) >DroEre_CAF1 7665 88 + 1 UUUGGGCAUUCGCCGGGCAAAAAUAAAUUGCUUGCAGCGG-GCGGCCAACUGGGUAAUGUGGCAACUAGGCAACC-------------------------AGCAACCAGGCAUC .((((.(.((((((((((((.......)))))))..))))-).).))))((((....((..((......))...)-------------------------)....))))..... ( -26.10) >DroYak_CAF1 7788 98 + 1 UUCAGACAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUUGGUAAUGUGGCAACUAGGCAACUUGGCACCUA----------------GGCAACCAGGCAUC ..........((((((((((.......))))((((....(((((((((.(........)))))..(.((....)).).))))).----------------.)))))).)))).. ( -28.10) >consensus UUCCGGCAUUUGCCGGGCAAAAAUAAAUUGCUUGCGGCGGGGUGGCCAACUGGGUAAUGUGGCAACUAGGCAACUGGGCA_CUAG________GCAACC_GGCAACCAGGCAUC ....(((.((((((((((((.......)))))..)))))))...)))..((((.......(....).((....))..............................))))..... (-20.84 = -21.60 + 0.76)


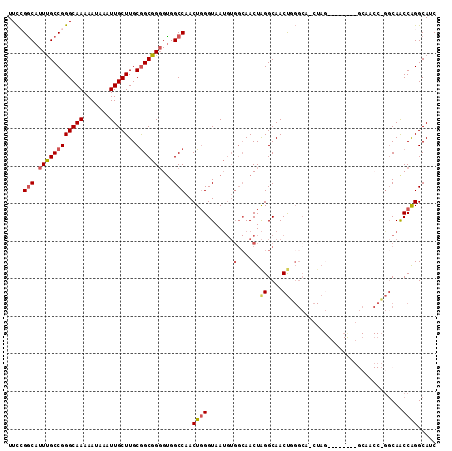
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:48:27 2006