
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,039,114 – 20,039,301 |
| Length | 187 |
| Max. P | 0.924113 |

| Location | 20,039,114 – 20,039,233 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.67 |
| Mean single sequence MFE | -36.35 |
| Consensus MFE | -24.47 |
| Energy contribution | -27.80 |
| Covariance contribution | 3.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.67 |
| SVM decision value | 1.16 |
| SVM RNA-class probability | 0.924113 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
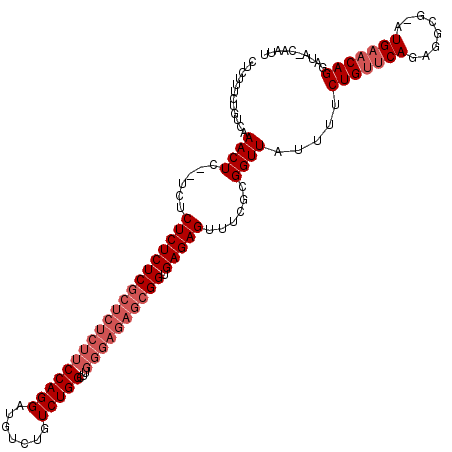
>3L_DroMel_CAF1 20039114 119 - 23771897 CUCUUUCUGUCAAACUCGCUCGCUCUCUCUCUCUCUUCCAGGAUGUCUGUCUGGCUUGGGAGAGCGGUGAGAGUUUCGCGGUUAUU-UCUGUUCAGAGGCGAAUGAACAGGAUAACAAUU ................(((..((((((.(.(.((((((((((..(.....)...)))))))))).)).))))))...)))((((((-.(((((((........))))))))))))).... ( -41.90) >DroSec_CAF1 6193 103 - 1 CUCUUUCUGUCAAACUC----UCUCUCUCGCUCUCUUCCAGGAUGUCUGUCUGGCUUGGGAGAGCGGUGAGAGUUUCGCGGUUAUUUUCUGUUCAGA------UGUUCAG-------AUU ......((((.((((((----((...(.((((((((((((((.......)))))...)))))))))).)))))))).))))......((((..(...------.)..)))-------).. ( -36.90) >DroSim_CAF1 7159 114 - 1 CUCUUUCUGUCAAACUCUUUCUCUCUCUCGCUCUCUUCCAGGAUGUCUGUCUGGCUC------UCGGUGAGAGUUUCGCGGUUAUUUUCUGUUCAGAGGCGGAUGAACAGGAUAGCAAUG ......(((((.((((........((((((((.....(((((.......)))))...------..))))))))......)))).....(((((((........))))))))))))..... ( -30.24) >consensus CUCUUUCUGUCAAACUC__UCUCUCUCUCGCUCUCUUCCAGGAUGUCUGUCUGGCUUGGGAGAGCGGUGAGAGUUUCGCGGUUAUUUUCUGUUCAGAGGCG_AUGAACAGGAUA_CAAUU ............((((......((((((((((((((((((((.......)))))...)))))))))).)))))......)))).....(((((((........))))))).......... (-24.47 = -27.80 + 3.33)


| Location | 20,039,193 – 20,039,301 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 91.44 |
| Mean single sequence MFE | -20.63 |
| Consensus MFE | -18.22 |
| Energy contribution | -18.56 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.88 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.894090 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 20039193 108 - 23771897 ACGUCAGUGUGACGUCAUGGUCGUGACGUUCGCACUCAAACUCUUUACCACCGAGGUUUU-UUACAUUCCUCUUUCUGUCAAACUCGCUCGCUCUCUCUCUCUCUUCCA .....(((((((((((((....)))))).)))))))...............(((((..((-(.(((..........))).)))..).)))).................. ( -21.60) >DroSec_CAF1 6260 105 - 1 ACGUCAGUGUGACGUCAUGGUCGUGACGUUCGCACUCAAACUCUUUACCAUCGACGUUCUUCUCGAUUCCUCUUUCUGUCAAACUC----UCUCUCUCGCUCUCUUCCA ((((((((((((((((((....)))))).)))))))................))))).......(((..........)))......----................... ( -20.19) >DroSim_CAF1 7233 109 - 1 ACGUCAGUGUGACGUCAUGGUCGUGACGUUCGCACUCAAACUCUUUACCAUCGACGUUCUUUUCCAUUCCUCUUUCUGUCAAACUCUUUCUCUCUCUCGCUCUCUUCCA ((((((((((((((((((....)))))).)))))))................))))).................................................... ( -20.09) >consensus ACGUCAGUGUGACGUCAUGGUCGUGACGUUCGCACUCAAACUCUUUACCAUCGACGUUCUUUUCCAUUCCUCUUUCUGUCAAACUC__UCUCUCUCUCGCUCUCUUCCA ((((((((((((((((((....)))))).)))))))................))))).................................................... (-18.22 = -18.56 + 0.33)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:48:14 2006