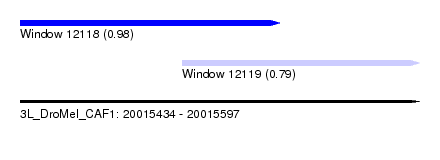
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,015,434 – 20,015,597 |
| Length | 163 |
| Max. P | 0.980369 |

| Location | 20,015,434 – 20,015,540 |
|---|---|
| Length | 106 |
| Sequences | 3 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 95.58 |
| Mean single sequence MFE | -16.83 |
| Consensus MFE | -17.05 |
| Energy contribution | -16.83 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.31 |
| Structure conservation index | 1.01 |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.980369 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
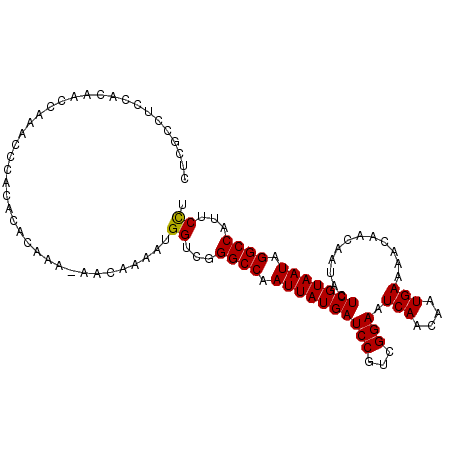
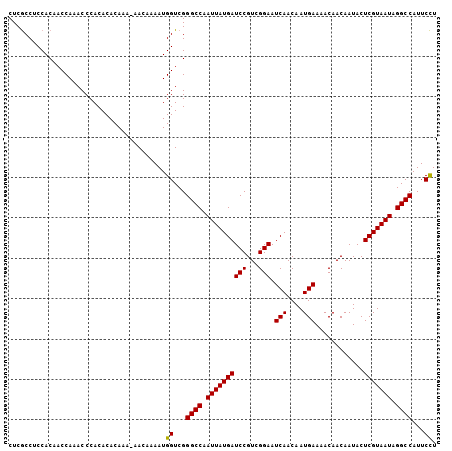
>3L_DroMel_CAF1 20015434 106 + 23771897 CUCACCCCCACACCCACACCCACACACAAAAAACAAAAUGGUCGGGCCAAUUAUGAUCCGUCGGAAUCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCUU ............................................((((.((((((((((...))).(((....)))............))))))).))))...... ( -15.10) >DroSec_CAF1 7403 105 + 1 CUCGCCUCCACCACCAAACCCACACACAAA-AACAAAAUGGUCGGGCCAAUUAUGAUCCGUCGGAAUCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCCU ..............................-........((...((((.((((((((((...))).(((....)))............))))))).))))...)). ( -17.70) >DroSim_CAF1 1209 105 + 1 CUCGCCUCCACAACCAAACCCACACACAAA-AACAAAAUGGUCGGGCCAAUUAUGAUCCGUCGGAAUCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCCU ..............................-........((...((((.((((((((((...))).(((....)))............))))))).))))...)). ( -17.70) >consensus CUCGCCUCCACAACCAAACCCACACACAAA_AACAAAAUGGUCGGGCCAAUUAUGAUCCGUCGGAAUCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCCU .......................................((...((((.((((((((((...))).(((....)))............))))))).))))...)). (-17.05 = -16.83 + -0.22)

| Location | 20,015,500 – 20,015,597 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 89.14 |
| Mean single sequence MFE | -22.70 |
| Consensus MFE | -17.72 |
| Energy contribution | -17.96 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.791923 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
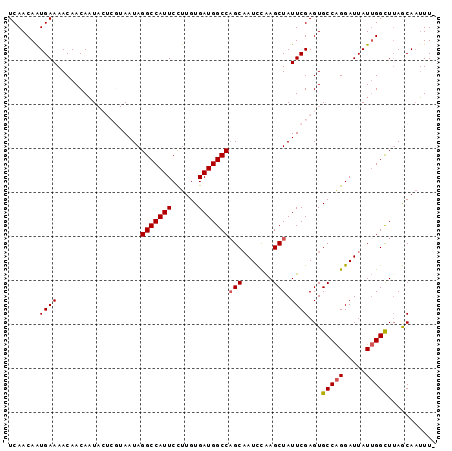
>3L_DroMel_CAF1 20015500 97 + 23771897 UCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCUUUGUGAUGGCCAGCAAUCUAAGCUAUUCGAGUGCCAGGAUUAUUGGCUUAGCAAUUUU ....................(((((.(((((((((((......))))))).((.......)))))))))))(((((.....)))))........... ( -24.50) >DroSec_CAF1 7468 96 + 1 UCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCCUUGUGAUGGCCAGCAAUCCAAGCUAUUCGAGUGCCAGGAUUAUUGGCUUAGCAAUUU- ....................(((((.(((((((((((......))))))).((.......)))))))))))(((((.....)))))..........- ( -24.50) >DroSim_CAF1 1274 96 + 1 UCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCCUUGUGAUGGCCAGCAAUCCAAGCUAUUCGAGUGCCAGGAUUAUUGGCUUAGCAAUUU- ....................(((((.(((((((((((......))))))).((.......)))))))))))(((((.....)))))..........- ( -24.50) >DroEre_CAF1 7251 89 + 1 UCAACAGUGAAAACAACAAU------AAUAGGCCAUUCCUUGUGAUGGCCAGCAAUUCGAGCUAUUCGAGUGCCGGGAUUAUGGGCUUAACA-UUU- ......((....))......------....(((((((......)))))))....(((((((...)))))))(((.(.....).)))......-...- ( -20.70) >DroYak_CAF1 8828 90 + 1 UCAACAAUGAAAACAACAUU------AAUAGGCCAUUCCUUGUGAUGGCCUGCCAUUCAAGCUAUUCGAGUGCCAGGAUUAUUGGUUAAACAAGUU- ....((((((......((((------(..(((.....)))..))))).((((.(((((.........))))).)))).))))))............- ( -19.30) >consensus UCAACAAUGAAAACAACAAUACUCGUAAUAGGCCAUUCCUUGUGAUGGCCAGCAAUCCAAGCUAUUCGAGUGCCAGGAUUAUUGGCUUAGCAAUUU_ .......((((...................(((((((......)))))))(((.......))).))))...(((((.....)))))........... (-17.72 = -17.96 + 0.24)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:47:36 2006