
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 20,011,225 – 20,011,350 |
| Length | 125 |
| Max. P | 0.898780 |

| Location | 20,011,225 – 20,011,338 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 85.97 |
| Mean single sequence MFE | -33.44 |
| Consensus MFE | -21.09 |
| Energy contribution | -21.03 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.689196 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
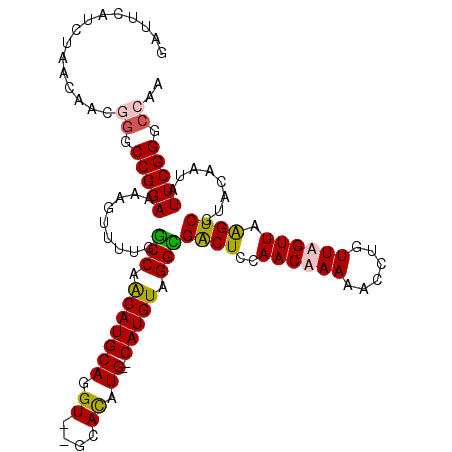
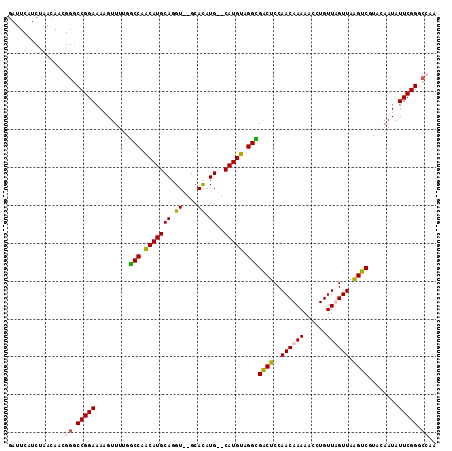
>3L_DroMel_CAF1 20011225 113 + 23771897 GAUUCAUCUAACAACGUUCCGGAAAAGUUUUGGCCAACAUGCAGGUGUGCACAUGCACAUGUAGGCGACUCCAACAAAAACCUGUUUGUUAAGUCGUCCAAUAUUCGGUCCAA ...............(..(((((.......((((.((((.((((((((((....)))).((.((....)).))......)))))).))))..)))).......)))))..).. ( -29.14) >DroSec_CAF1 3235 111 + 1 GAUUCAUCUAACAACGGGCCGGAAAAGUUUUGGCCAACAUGCAGGU--GCACAUGCACAUGUUGGCGACUCCAACAAAAACCUGUUUGUUAGGUCGUCCAAUAUUCGGGCGAA ......((((((((..(((((((.....))))))).....((((((--((....))...((((((.....))))))...))))))))))))))((((((.......)))))). ( -40.10) >DroEre_CAF1 3325 107 + 1 GAUUCAGCCGACAAAGGGCCGGAAAAGUUU-GGCCAACAUGCAGGU--GCAU-UG--CAUGUAGGUGGCUCCAACAAAAAGCUGUUAGUUAAGUCGUAGAAUAUUCGGGCCGA ......(((.......(((((((....)))-)))).((((((((..--...)-))--))))).)))(((.((((((......))))............((....))))))).. ( -31.00) >DroYak_CAF1 4145 109 + 1 GAUUCAUCUAACAACGGGCCGGAAAAGUUUUGCCCAGCAUGCAGGU--GGAUAUG--CAUGUAGGGGACUCCAACAAAAAGCUGUUAGUUAAGUCGUAGAAUAUUCGGCCAAA ................(((((((...(((((((((.(((((((...--.....))--))))).)).(((((.((((......)))).)...))))))))))).)))))))... ( -33.50) >consensus GAUUCAUCUAACAACGGGCCGGAAAAGUUUUGGCCAACAUGCAGGU__GCACAUG__CAUGUAGGCGACUCCAACAAAAACCUGUUAGUUAAGUCGUACAAUAUUCGGGCCAA ...............((.(((((.........(((.(((((((.((....)).))..))))).)))((((..((((((......)))))).))))........))))).)).. (-21.09 = -21.03 + -0.06)

| Location | 20,011,258 – 20,011,350 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 86.58 |
| Mean single sequence MFE | -26.32 |
| Consensus MFE | -18.11 |
| Energy contribution | -17.80 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.69 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.898780 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
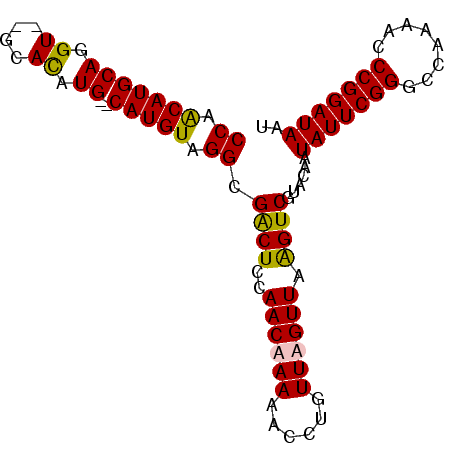
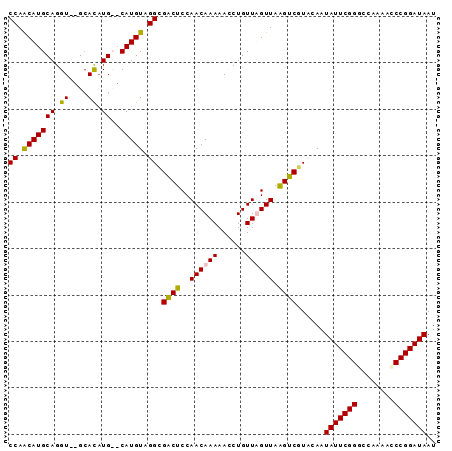
>3L_DroMel_CAF1 20011258 92 + 23771897 CCAACAUGCAGGUGUGCACAUGCACAUGUAGGCGACUCCAACAAAAACCUGUUUGUUAAGUCGUCCAAUAUUCGGUCCAAAACCCGGAUAAU ......((((..(((((....)))))))))(((((((..((((((......)))))).)))))))...(((((((........))))))).. ( -28.10) >DroSec_CAF1 3268 90 + 1 CCAACAUGCAGGU--GCACAUGCACAUGUUGGCGACUCCAACAAAAACCUGUUUGUUAGGUCGUCCAAUAUUCGGGCGAAAACCCGGAUAAU ((((((.((((((--((....))...((((((.....))))))...)))))).)))).))........((((((((......)))))))).. ( -31.70) >DroEre_CAF1 3357 87 + 1 CCAACAUGCAGGU--GCAU-UG--CAUGUAGGUGGCUCCAACAAAAAGCUGUUAGUUAAGUCGUAGAAUAUUCGGGCCGAAAUCCGGAUAAU ((.((((((((..--...)-))--))))).)).(((((.((((......)))).)...))))......((((((((......)))))))).. ( -22.70) >DroYak_CAF1 4178 88 + 1 CCAGCAUGCAGGU--GGAUAUG--CAUGUAGGGGACUCCAACAAAAAGCUGUUAGUUAAGUCGUAGAAUAUUCGGCCAAAAACCCGGAUAAU ((.(((((((...--.....))--))))).)).(((((.((((......)))).)...))))......(((((((........))))))).. ( -22.80) >consensus CCAACAUGCAGGU__GCACAUG__CAUGUAGGCGACUCCAACAAAAACCUGUUAGUUAAGUCGUACAAUAUUCGGGCCAAAACCCGGAUAAU ((.(((((((.((....)).))..))))).)).((((..((((((......)))))).))))......(((((((........))))))).. (-18.11 = -17.80 + -0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:47:31 2006