| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,980,292 – 19,980,409 |
| Length | 117 |
| Max. P | 0.743756 |

| Location | 19,980,292 – 19,980,409 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.00 |
| Mean single sequence MFE | -28.64 |
| Consensus MFE | -12.16 |
| Energy contribution | -13.28 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.42 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.601571 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
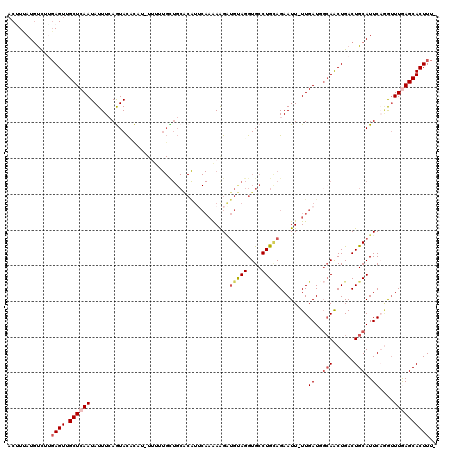
>3L_DroMel_CAF1 19980292 117 + 23771897 ACUUUAUGUUUUGAGUUGCUCAAUAUUUCAGUACACUU-UUUAUGUUGCGCAUUCAAAAAGAUGCAGGUGCCUGCAGAAUU-UUGAUGGCAACUGACUGCAUUCAGGUUUGAGCACUUU- ............((((.((((((...............-.....(((((.(((.(((((...(((((....)))))...))-)))))))))))...(((....)))..)))))))))).- ( -31.80) >DroSec_CAF1 70586 118 + 1 ACAUUAUGUCUUGAGUUGCUCAAUAUUUCAGUACUCAUAUAUUUGCUGCACAUUCAAAAAGAUGUAGGUGCCUGCAGAAUU-UUGAUGGCAACUGACUGCAUUCCGGUUUGAGCACUUU- ............((((.((((((....(((((...((......)).(((.(((.(((((...(((((....)))))...))-))))))))))))))(((.....))).)))))))))).- ( -27.90) >DroSim_CAF1 29238 115 + 1 ACUUUAUGUCUUGAGUUGCUCAAUAUUUCAGUACUCAUAUUUUUGCUGCACAUUCAAAAAGGUGUAGGUGCCUGUAGAAUU-UUGAU---AACUGACUGCAUUCCAGUUUGAGCACUUU- ............((((.((((((....(((((..(((...((((((.((((...((......))...))))..))))))..-.))).---.)))))(((.....))).)))))))))).- ( -28.60) >DroEre_CAF1 21134 119 + 1 ACCUUAUCCCUGCAGUUGCUCAAUAUUUGAAUACCUUAUUUUGUGUUGCACGUGCAAGAGGAUGUAGGUGCCUGCAGAAUU-UUGAUUGCAACUGACUGCAUUCAGAAUUAAGCACUUUU ((((((((((((((..(((..(((((..(((((...))))).))))))))..))).)).))))).))))...(((..((((-((((.((((......)))).))))))))..)))..... ( -34.10) >DroYak_CAF1 22040 97 + 1 -CUUCAUGUCUGGAGUUGCUCAAUAUUUAAAUAUAU---------------------AAGGUGGUAGGUGCCUGAAGAAUUUUUGAUUGCAGCUGACUGCAUUCACAUUUAAGCACUUU- -(((((....)))))(..((..((((......))))---------------------..))..).((((((.((((.......(((.(((((....))))).)))..)))).)))))).- ( -20.80) >consensus ACUUUAUGUCUUGAGUUGCUCAAUAUUUCAGUACACAU_UUUUUGCUGCACAUUCAAAAAGAUGUAGGUGCCUGCAGAAUU_UUGAUGGCAACUGACUGCAUUCAGGUUUGAGCACUUU_ ............((((.((((((.......................................(((((....)))))........((..(((......)))..))....)))))))))).. (-12.16 = -13.28 + 1.12)


| Location | 19,980,292 – 19,980,409 |
|---|---|
| Length | 117 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 77.00 |
| Mean single sequence MFE | -25.06 |
| Consensus MFE | -10.96 |
| Energy contribution | -12.92 |
| Covariance contribution | 1.96 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.44 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.743756 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
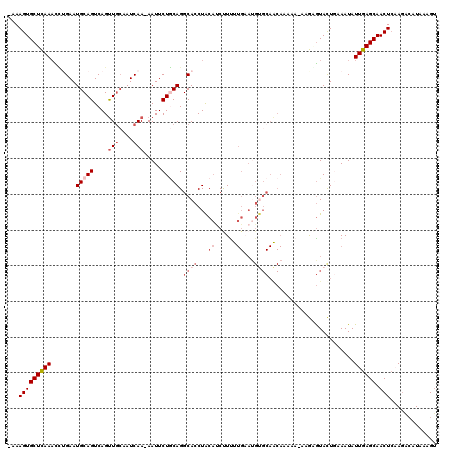
>3L_DroMel_CAF1 19980292 117 - 23771897 -AAAGUGCUCAAACCUGAAUGCAGUCAGUUGCCAUCAA-AAUUCUGCAGGCACCUGCAUCUUUUUGAAUGCGCAACAUAAA-AAGUGUACUGAAAUAUUGAGCAACUCAAAACAUAAAGU -....(((((((..(((....)))((((((((((((((-((...(((((....)))))...))))).))).)))((((...-..)))))))))....)))))))................ ( -32.30) >DroSec_CAF1 70586 118 - 1 -AAAGUGCUCAAACCGGAAUGCAGUCAGUUGCCAUCAA-AAUUCUGCAGGCACCUACAUCUUUUUGAAUGUGCAGCAAAUAUAUGAGUACUGAAAUAUUGAGCAACUCAAGACAUAAUGU -..((((((((.....((......)).(((((((((((-((...((.((....)).))...))))).))).))))).......))))))))......(((((...))))).......... ( -26.60) >DroSim_CAF1 29238 115 - 1 -AAAGUGCUCAAACUGGAAUGCAGUCAGUU---AUCAA-AAUUCUACAGGCACCUACACCUUUUUGAAUGUGCAGCAAAAAUAUGAGUACUGAAAUAUUGAGCAACUCAAGACAUAAAGU -..((((((((((((((.......))))))---.....-..........((((...((......))...))))..........))))))))......(((((...))))).......... ( -21.30) >DroEre_CAF1 21134 119 - 1 AAAAGUGCUUAAUUCUGAAUGCAGUCAGUUGCAAUCAA-AAUUCUGCAGGCACCUACAUCCUCUUGCACGUGCAACACAAAAUAAGGUAUUCAAAUAUUGAGCAACUGCAGGGAUAAGGU .....(((..((((.(((.(((((....))))).))).-))))..)))...((((..(((((..((((..(((...((........))..(((.....))))))..))))))))).)))) ( -27.30) >DroYak_CAF1 22040 97 - 1 -AAAGUGCUUAAAUGUGAAUGCAGUCAGCUGCAAUCAAAAAUUCUUCAGGCACCUACCACCUU---------------------AUAUAUUUAAAUAUUGAGCAACUCCAGACAUGAAG- -....(((((((...(((.(((((....))))).)))...........((......)).....---------------------.............)))))))...............- ( -17.80) >consensus _AAAGUGCUCAAACCUGAAUGCAGUCAGUUGCAAUCAA_AAUUCUGCAGGCACCUACAUCUUUUUGAAUGUGCAACAAAAA_AAGAGUACUGAAAUAUUGAGCAACUCAAGACAUAAAGU ...(((((((((.......(((((....(((....))).....))))).((((...((......))...))))........................)))))).)))............. (-10.96 = -12.92 + 1.96)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:46:47 2006