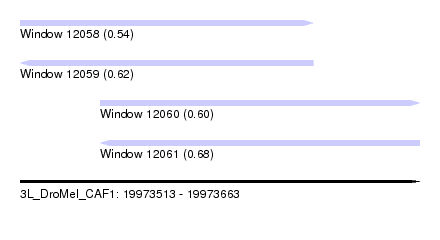

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,973,513 – 19,973,663 |
| Length | 150 |
| Max. P | 0.680828 |

| Location | 19,973,513 – 19,973,623 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 84.00 |
| Mean single sequence MFE | -32.36 |
| Consensus MFE | -23.52 |
| Energy contribution | -24.64 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.537646 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19973513 110 + 23771897 CGCACGCUCCAUAUUGUAGUGAUGUUAAUUACUUAUUCUCUCGGUGAACAUGUGACAAACUCACUGGAGUUCUUUCAGCUCCAAGUGAGCUGGUGUUUGUGGGCAGAUAA ((((.((.(((..((((((((((....))))))..(((.......)))......)))).((((((((((((.....)))))).)))))).))).)).))))......... ( -29.70) >DroSec_CAF1 64369 110 + 1 CGCACGCUCCAUAUUGUAGUGAUGUUAAUUACUUAUUCUCUCGGUGAGCAUGUGACAAACUCACUGGAGUUCUUUCAACUCCAAGUGAGCUGGUGCUGGUGGGCAGAUAA ((((.(((((.....(.((((((....))))))....).....).)))).))))((...((((((((((((.....)))))).))))))...))(((....)))...... ( -32.00) >DroSim_CAF1 22952 110 + 1 CGCACGCUCCAUAUUGUAGUGAUGUUAAUUACUUAUUCUCUCGGUGAGCAUGUGACAAACUCACUGGAGUUCUUUCAGCUCCAAGUGAGCUGGUGCUUGUGGGCAGAUAA ((((.((.(((..((((((((((....))))))(((.(((.....))).)))..)))).((((((((((((.....)))))).)))))).))).)).))))......... ( -33.70) >DroEre_CAF1 15282 94 + 1 CGCACGCU---------------GUUAAUAACUUAUUCGCUGGGUGAGCGUGUGACCAACUCACUGGAGUUCUGUCUGCUCCAAGUGAGCUUGUUUCUAUGCGAAUAG-A ((((((((---------------.......(((((.....))))).))))))))..((((((((((((((.......))))).)))))).)))..(((((....))))-) ( -31.20) >DroYak_CAF1 15573 110 + 1 CGCACGCUCCGUAGAGUAGUGACGUUAAUUACUUAUUCUCCGGGAGAGCAUGUGACAAACUCGCUGGAGUUCUUUCUGCUCCAAGUGAGCUGGUGUUUAUGCGAAGAUAA ((((.((((((.((((((((((......)))).)))))).)))...)))....((((..(((((((((((.......))))).))))))....))))..))))....... ( -35.20) >consensus CGCACGCUCCAUAUUGUAGUGAUGUUAAUUACUUAUUCUCUCGGUGAGCAUGUGACAAACUCACUGGAGUUCUUUCAGCUCCAAGUGAGCUGGUGUUUGUGGGCAGAUAA ((((.((.(((............((((....((((((.....))))))....))))...(((((((((((.......))))).)))))).))).)).))))......... (-23.52 = -24.64 + 1.12)



| Location | 19,973,513 – 19,973,623 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 84.00 |
| Mean single sequence MFE | -28.08 |
| Consensus MFE | -22.88 |
| Energy contribution | -22.96 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.58 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.623317 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19973513 110 - 23771897 UUAUCUGCCCACAAACACCAGCUCACUUGGAGCUGAAAGAACUCCAGUGAGUUUGUCACAUGUUCACCGAGAGAAUAAGUAAUUAACAUCACUACAAUAUGGAGCGUGCG ......(((((.......((((((.....))))))..........((((((((.((.((.(((((.......))))).)).)).))).)))))......))).))..... ( -26.10) >DroSec_CAF1 64369 110 - 1 UUAUCUGCCCACCAGCACCAGCUCACUUGGAGUUGAAAGAACUCCAGUGAGUUUGUCACAUGCUCACCGAGAGAAUAAGUAAUUAACAUCACUACAAUAUGGAGCGUGCG ..............((((.((((((((.((((((.....))))))))))))))........((((..(....)..(((....)))................)))))))). ( -30.80) >DroSim_CAF1 22952 110 - 1 UUAUCUGCCCACAAGCACCAGCUCACUUGGAGCUGAAAGAACUCCAGUGAGUUUGUCACAUGCUCACCGAGAGAAUAAGUAAUUAACAUCACUACAAUAUGGAGCGUGCG ..............((((((((((.....))))))......(((((((((((.((...)).))))))...((...(((....)))...)).........))))).)))). ( -30.40) >DroEre_CAF1 15282 94 - 1 U-CUAUUCGCAUAGAAACAAGCUCACUUGGAGCAGACAGAACUCCAGUGAGUUGGUCACACGCUCACCCAGCGAAUAAGUUAUUAAC---------------AGCGUGCG .-.....(((((.((....((((((((.((((.........))))))))))))..))...((((.....))))..............---------------...))))) ( -26.20) >DroYak_CAF1 15573 110 - 1 UUAUCUUCGCAUAAACACCAGCUCACUUGGAGCAGAAAGAACUCCAGCGAGUUUGUCACAUGCUCUCCCGGAGAAUAAGUAAUUAACGUCACUACUCUACGGAGCGUGCG .......(((((.........(((....((((((....((((((....))))))......))))))....))).....................(((....))).))))) ( -26.90) >consensus UUAUCUGCCCACAAACACCAGCUCACUUGGAGCUGAAAGAACUCCAGUGAGUUUGUCACAUGCUCACCGAGAGAAUAAGUAAUUAACAUCACUACAAUAUGGAGCGUGCG ................((.((((((((.((((.........)))))))))))).))..(((((((..(....)..(((....)))................))))))).. (-22.88 = -22.96 + 0.08)


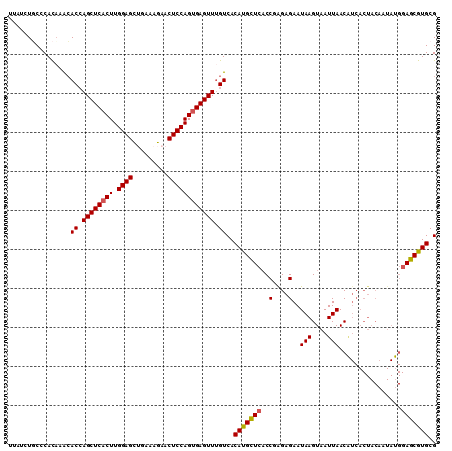
| Location | 19,973,543 – 19,973,663 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 86.33 |
| Mean single sequence MFE | -36.38 |
| Consensus MFE | -24.48 |
| Energy contribution | -24.56 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.597251 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19973543 120 + 23771897 ACUUAUUCUCUCGGUGAACAUGUGACAAACUCACUGGAGUUCUUUCAGCUCCAAGUGAGCUGGUGUUUGUGGGCAGAUAAUCUGCCAAAUCAUGUCCUGGAUUUUUGCUCAAGUUUCAGC ............(.((((....(((((((((((((((((((.....)))))).))))))(..(..(.((..(((((.....)))))....)).)..)..)...)))).)))...)))).) ( -34.20) >DroSec_CAF1 64399 120 + 1 ACUUAUUCUCUCGGUGAGCAUGUGACAAACUCACUGGAGUUCUUUCAACUCCAAGUGAGCUGGUGCUGGUGGGCAGAUAAUCUGCCAAAUCAUGUCCUGGAUUUUUGCUCAGGUUUCAGC ..............((((((.........((((((((((((.....)))))).))))))(..(..(((((.(((((.....)))))..)))).)..)..).....))))))......... ( -39.40) >DroSim_CAF1 22982 120 + 1 ACUUAUUCUCUCGGUGAGCAUGUGACAAACUCACUGGAGUUCUUUCAGCUCCAAGUGAGCUGGUGCUUGUGGGCAGAUAAUCUGCCAAAUCAUGCACUGGAUUUUUGCUCAAGUUUCAGA ......(((.....((((((.........((((((((((((.....)))))).))))))(..((((.((..(((((.....)))))....)).))))..).....))))))......))) ( -44.50) >DroEre_CAF1 15297 118 + 1 ACUUAUUCGCUGGGUGAGCGUGUGACCAACUCACUGGAGUUCUGUCUGCUCCAAGUGAGCUUGUUUCUAUGCGAAUAG-AUUUGCCUAAUAAUGG-CUGGAUUUUCCCUUAAGUUACAGC ........((((.....(((((.((.((((((((((((((.......))))).)))))).))).)).)))))(((..(-(((((((.......))-).))))))))..........)))) ( -33.70) >DroYak_CAF1 15603 120 + 1 ACUUAUUCUCCGGGAGAGCAUGUGACAAACUCGCUGGAGUUCUUUCUGCUCCAAGUGAGCUGGUGUUUAUGCGAAGAUAAUCUGCCAAAUCAUGUCCUAGAUUUUUGCUUGAGUUUUAGC .((....(((.((.((((((((.((((..(((((((((((.......))))).))))))....)))))))))........))).))(((((........)))))......)))....)). ( -30.10) >consensus ACUUAUUCUCUCGGUGAGCAUGUGACAAACUCACUGGAGUUCUUUCAGCUCCAAGUGAGCUGGUGUUUGUGGGCAGAUAAUCUGCCAAAUCAUGUCCUGGAUUUUUGCUCAAGUUUCAGC ..............((((((.........(((((((((((.......))))).))))))((((.((.....(((((.....))))).......)).)))).....))))))......... (-24.48 = -24.56 + 0.08)


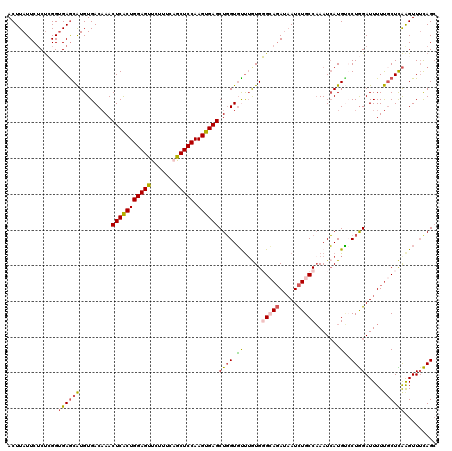
| Location | 19,973,543 – 19,973,663 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.33 |
| Mean single sequence MFE | -33.74 |
| Consensus MFE | -21.86 |
| Energy contribution | -23.38 |
| Covariance contribution | 1.52 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.680828 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19973543 120 - 23771897 GCUGAAACUUGAGCAAAAAUCCAGGACAUGAUUUGGCAGAUUAUCUGCCCACAAACACCAGCUCACUUGGAGCUGAAAGAACUCCAGUGAGUUUGUCACAUGUUCACCGAGAGAAUAAGU (((........))).........(((((((....(((((.....))))).......((.((((((((.((((.........)))))))))))).))..)))))))............... ( -35.50) >DroSec_CAF1 64399 120 - 1 GCUGAAACCUGAGCAAAAAUCCAGGACAUGAUUUGGCAGAUUAUCUGCCCACCAGCACCAGCUCACUUGGAGUUGAAAGAACUCCAGUGAGUUUGUCACAUGCUCACCGAGAGAAUAAGU ((((...((((..........)))).........(((((.....)))))...))))((.((((((((.((((((.....)))))))))))))).))......(((.....)))....... ( -39.40) >DroSim_CAF1 22982 120 - 1 UCUGAAACUUGAGCAAAAAUCCAGUGCAUGAUUUGGCAGAUUAUCUGCCCACAAGCACCAGCUCACUUGGAGCUGAAAGAACUCCAGUGAGUUUGUCACAUGCUCACCGAGAGAAUAAGU ......(((((((((........((((.((....(((((.....)))))..)).)))).((((((((.((((.........)))))))))))).......)))))..(....)...)))) ( -37.30) >DroEre_CAF1 15297 118 - 1 GCUGUAACUUAAGGGAAAAUCCAG-CCAUUAUUAGGCAAAU-CUAUUCGCAUAGAAACAAGCUCACUUGGAGCAGACAGAACUCCAGUGAGUUGGUCACACGCUCACCCAGCGAAUAAGU .............((.....)).(-((.......)))....-.(((((((.........((((((((.((((.........))))))))))))(((.........)))..)))))))... ( -30.20) >DroYak_CAF1 15603 120 - 1 GCUAAAACUCAAGCAAAAAUCUAGGACAUGAUUUGGCAGAUUAUCUUCGCAUAAACACCAGCUCACUUGGAGCAGAAAGAACUCCAGCGAGUUUGUCACAUGCUCUCCCGGAGAAUAAGU (((........))).....((..(((((((...((((((((.....((((..........((((.....)))).((......))..))))))))))))))))...)))..))........ ( -26.30) >consensus GCUGAAACUUGAGCAAAAAUCCAGGACAUGAUUUGGCAGAUUAUCUGCCCACAAACACCAGCUCACUUGGAGCUGAAAGAACUCCAGUGAGUUUGUCACAUGCUCACCGAGAGAAUAAGU .........((((((...................(((((.....))))).......((.((((((((.((((.........)))))))))))).))....)))))).............. (-21.86 = -23.38 + 1.52)


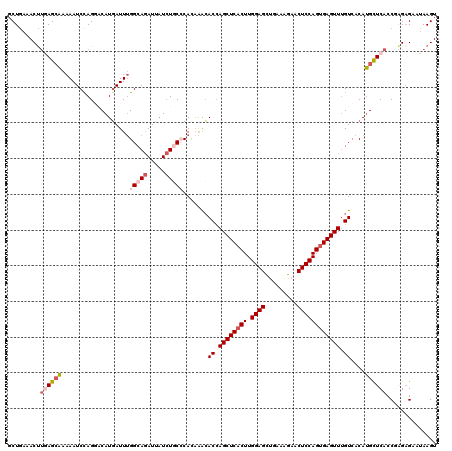
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:46:41 2006