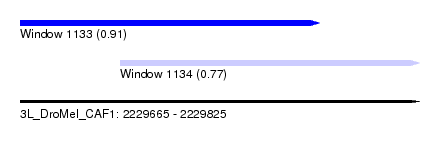
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 2,229,665 – 2,229,825 |
| Length | 160 |
| Max. P | 0.909925 |

| Location | 2,229,665 – 2,229,785 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 91.58 |
| Mean single sequence MFE | -38.06 |
| Consensus MFE | -29.66 |
| Energy contribution | -30.06 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.07 |
| SVM RNA-class probability | 0.909925 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
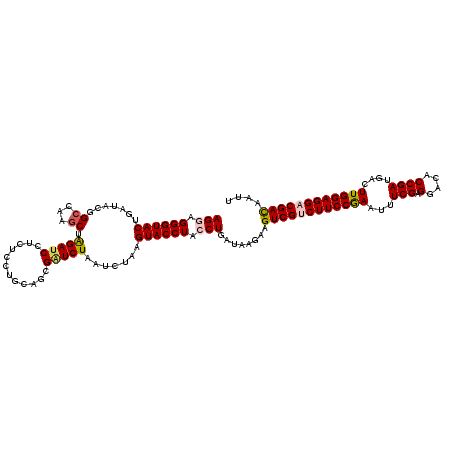
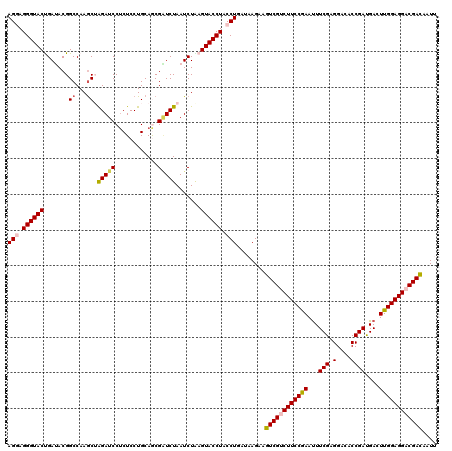
>3L_DroMel_CAF1 2229665 120 + 23771897 AGGAGGGUACUGAUACGGCCAAGCUGGAUCCUCUCCUGCAGCGGUCUAAUCUGAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACACCGACGACUUGGAGGACGAUAAUU (((.((((((((((..((((..((.(((.....))).))...))))..)))..))))))).)))........((((((((((((.(((((.(....)))).)).)))))))))))).... ( -43.50) >DroSec_CAF1 10595 120 + 1 AGAAGGGUACUGAUACGGCCAAGCUAGAUCCUCUCCUGCAGCGAUCUAAUCUAAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACGCCGAUGACUUGGAGGACGACAAUU ....((((((((((..(((...)))((((((((....).)).))))).)))..)))))))............((((((((((((...(((.......)))....)))))))))))).... ( -36.70) >DroSim_CAF1 10684 120 + 1 AGAAGGGUACUGAUACGGCCAAGCUAGAUCCUCUCCUGCAGCGAUCUAAUCUAAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACGCCGAUGACUUGGAGGACGACAAUU ....((((((((((..(((...)))((((((((....).)).))))).)))..)))))))............((((((((((((...(((.......)))....)))))))))))).... ( -36.70) >DroEre_CAF1 23620 119 + 1 AGGAGGGUACCGAAACGGCCAAGCAAGAUCC-CUCCUGCAACGGUCUAAUCUGAGUACCUACCUGAUAAGAAGUCGACUUCCAAACUUCGAGGACACCGAUGACUUGGAGGACGACAAUA (((.(((((((((...((((..(((.((...-.)).)))...))))...)).).)))))).)))........((((.(((((((...(((.(....))))....))))))).)))).... ( -37.10) >DroYak_CAF1 11117 120 + 1 AGGAGGGUACCGAGACGGCCAAACAGGAUCCUCUCCUGCAACGUUCUAAUCUAAGUACCUACCUAAUAAGAAGUCGACUUCCAAAUUUCGAGGACUCCGAUGACUUGGAGGACGACAAUA (((.((((((..(((.(((....(((((.....)))))....)))....)))..)))))).)))........((((.(((((((...(((.......)))....))))))).)))).... ( -36.30) >consensus AGGAGGGUACUGAUACGGCCAAGCUAGAUCCUCUCCUGCAGCGAUCUAAUCUAAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACACCGAUGACUUGGAGGACGACAAUU (((.((((((.......((...)).(((((............))))).......)))))).)))........((((((((((((...(((.(....))))....)))))))))))).... (-29.66 = -30.06 + 0.40)

| Location | 2,229,705 – 2,229,825 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 90.00 |
| Mean single sequence MFE | -32.24 |
| Consensus MFE | -27.50 |
| Energy contribution | -27.74 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.53 |
| SVM RNA-class probability | 0.771019 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
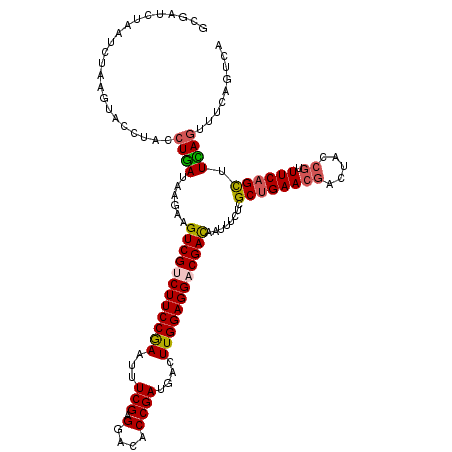
>3L_DroMel_CAF1 2229705 120 + 23771897 GCGGUCUAAUCUGAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACACCGACGACUUGGAGGACGAUAAUUUCUCGCUGAACGACUACCGUUUCAGCUUCAGUUUCAGUCA ..(((((......)).)))...((((...(((((((((((((((.(((((.(....)))).)).))))))))))))........((((((((.....)).)))))))))....))))... ( -35.80) >DroSec_CAF1 10635 120 + 1 GCGAUCUAAUCUAAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACGCCGAUGACUUGGAGGACGACAAUUUCUCGCUGAACGACUAUCGUUUCAGCUUCAGUUUUAGUGA .................((((.((((......((((((((((((...(((.......)))....))))))))))))........((((((((.....)).)))))).))))...))).). ( -34.70) >DroSim_CAF1 10724 120 + 1 GCGAUCUAAUCUAAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACGCCGAUGACUUGGAGGACGACAAUUUCCCGCUGAACGACUAUCGUUUCAGCUUCAGUUUCAGUGA .....................(((((...(((((((((((((((...(((.......)))....))))))))))))........((((((((.....)).)))))))))....)))).). ( -35.40) >DroEre_CAF1 23659 120 + 1 ACGGUCUAAUCUGAGUACCUACCUGAUAAGAAGUCGACUUCCAAACUUCGAGGACACCGAUGACUUGGAGGACGACAAUAUCUCGCUGAACGACUACCGUUUCAGCUUUAGUUUCAGUCA ..(((((......)).)))...((((......((((.(((((((...(((.(....))))....))))))).))))........((((((((.....)).)))))).......))))... ( -32.40) >DroYak_CAF1 11157 120 + 1 ACGUUCUAAUCUAAGUACCUACCUAAUAAGAAGUCGACUUCCAAAUUUCGAGGACUCCGAUGACUUGGAGGACGACAAUAUCUCGCAGAACGACUACAACUUCAGUUUUAUUUUCACCCA .((((((..(((...((......))...))).((((.(((((((...(((.......)))....))))))).))))..........))))))............................ ( -22.90) >consensus GCGAUCUAAUCUAAGUACCUACCUGAUAAGAAGUCGUCUUCCGAAUUUCGAGGACACCGAUGACUUGGAGGACGACAAUUUCUCGCUGAACGACUACCGUUUCAGCUUCAGUUUCAGUCA ......................((((......((((((((((((...(((.(....))))....))))))))))))........((((((((.....)).)))))).))))......... (-27.50 = -27.74 + 0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:49:55 2006