
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,627,539 – 19,627,638 |
| Length | 99 |
| Max. P | 0.998779 |

| Location | 19,627,539 – 19,627,638 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 87.12 |
| Mean single sequence MFE | -47.44 |
| Consensus MFE | -36.68 |
| Energy contribution | -39.36 |
| Covariance contribution | 2.68 |
| Combinations/Pair | 1.09 |
| Mean z-score | -4.28 |
| Structure conservation index | 0.77 |
| SVM decision value | 3.22 |
| SVM RNA-class probability | 0.998779 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
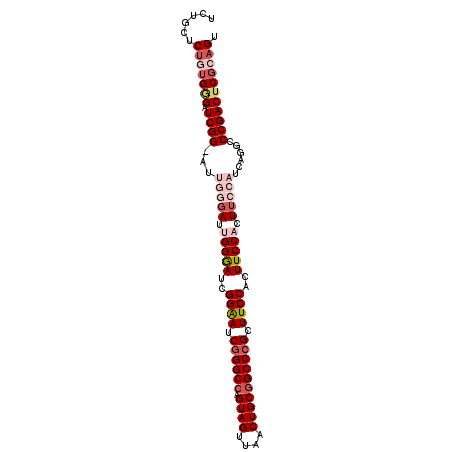
>3L_DroMel_CAF1 19627539 99 + 23771897 UCUGCUCUGUGGGAUCGGUAUUGGGACUGGAAUCGGAAUCGGGCCAGUAGUUAACUGCGGCCCGCUUCCACUUCCACU-CCAUCAGGCCCGACUCGCAGU ......(((((((.((((..((((((.(((((..((((.((((((.((((....)))))))))).))))..))))).)-))..)))..))))))))))). ( -50.40) >DroSec_CAF1 42284 100 + 1 UCUGCUCUGUGGGAUCGGGAUUGGGAUUGGGAUCGGAAUCGGGCCAGUAGUUAACUGCGGCCCGCUUCCACUUCCACUUCCAUCAGGCCCGACUCGCAGU ......(((((((.(((((..(((((.(((((..((((.((((((.((((....)))))))))).))))..))))).))))).....)))))))))))). ( -53.90) >DroSim_CAF1 44459 100 + 1 UCUGCUCUGUGUGAUCGGGAUUGGGAUUGGGAUCGGAAUCGGGCCAGUAGUUAACUGCGGCCCGCUUCCACUUCCACUUCCAUCAGGCCCGACUCGCAGU ......(((((.(.(((((..(((((.(((((..((((.((((((.((((....)))))))))).))))..))))).))))).....)))))).))))). ( -50.60) >DroEre_CAF1 42755 94 + 1 GUUAGGCAUAGGGAUCGG------GAUCGGGAUCGGGAUCCGGCCAGUAGUUAACUGCAGCCCGCUUCCACUUCCACUUCCAUCAGGCCCGACUCGCAGU ((..(((....(((...(------((..((((.((((.....((.(((.....)))))..)))).))))...)))...))).....)))..))....... ( -31.90) >DroYak_CAF1 42552 94 + 1 GCUGCUCUGUGAGAUCGG------GAUAGGGAUUGGGAUCGGGCCUGUAGUUAACUGCGGCCCGAUUCCACUUCCACUUCCAUCAGGCCCGACUCGGAGU ...(((((..(((.((((------(...((((.((((((((((((.((((....)))))))))))))))).))))....((....))))))))))))))) ( -50.40) >consensus UCUGCUCUGUGGGAUCGG_AUUGGGAUUGGGAUCGGAAUCGGGCCAGUAGUUAACUGCGGCCCGCUUCCACUUCCACUUCCAUCAGGCCCGACUCGCAGU ......(((((((.((((...(((((.(((((..((((.((((((.((((....)))))))))).))))..))))).)))))......))))))))))). (-36.68 = -39.36 + 2.68)



| Location | 19,627,539 – 19,627,638 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 87.12 |
| Mean single sequence MFE | -37.42 |
| Consensus MFE | -26.70 |
| Energy contribution | -27.42 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.680905 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19627539 99 - 23771897 ACUGCGAGUCGGGCCUGAUGG-AGUGGAAGUGGAAGCGGGCCGCAGUUAACUACUGGCCCGAUUCCGAUUCCAGUCCCAAUACCGAUCCCACAGAGCAGA .((((..(((((...((..((-(.(((((.(((((.((((((..(((.....))))))))).))))).))))).)))))...)))))........)))). ( -43.20) >DroSec_CAF1 42284 100 - 1 ACUGCGAGUCGGGCCUGAUGGAAGUGGAAGUGGAAGCGGGCCGCAGUUAACUACUGGCCCGAUUCCGAUCCCAAUCCCAAUCCCGAUCCCACAGAGCAGA .((((..((((((..((..(((..(((...(((((.((((((..(((.....))))))))).)))))...))).)))))..))))))........)))). ( -39.60) >DroSim_CAF1 44459 100 - 1 ACUGCGAGUCGGGCCUGAUGGAAGUGGAAGUGGAAGCGGGCCGCAGUUAACUACUGGCCCGAUUCCGAUCCCAAUCCCAAUCCCGAUCACACAGAGCAGA .((((...(((((..((..(((..(((...(((((.((((((..(((.....))))))))).)))))...))).)))))..)))))((.....)))))). ( -39.80) >DroEre_CAF1 42755 94 - 1 ACUGCGAGUCGGGCCUGAUGGAAGUGGAAGUGGAAGCGGGCUGCAGUUAACUACUGGCCGGAUCCCGAUCCCGAUC------CCGAUCCCUAUGCCUAAC .......((..(((.....(((.(.(((.(.(((..(((((((((((.....))))..)))..)))).))))..))------))..)))....)))..)) ( -25.90) >DroYak_CAF1 42552 94 - 1 ACUCCGAGUCGGGCCUGAUGGAAGUGGAAGUGGAAUCGGGCCGCAGUUAACUACAGGCCCGAUCCCAAUCCCUAUC------CCGAUCUCACAGAGCAGC .(((.((((((((((....)).((.(((..(((.((((((((..((....))...)))))))).))).)))))..)------)))).)))...))).... ( -38.60) >consensus ACUGCGAGUCGGGCCUGAUGGAAGUGGAAGUGGAAGCGGGCCGCAGUUAACUACUGGCCCGAUUCCGAUCCCAAUCCCAAU_CCGAUCCCACAGAGCAGA .(((.(.(((((.((....))..(.(((..(((((.(((((((.((....)).).)))))).))))).))))..........)))))..).)))...... (-26.70 = -27.42 + 0.72)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:44:03 2006