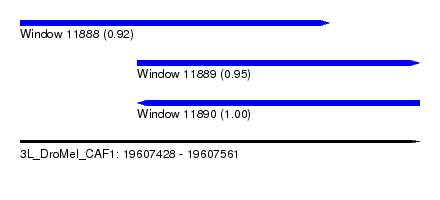

| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,607,428 – 19,607,561 |
| Length | 133 |
| Max. P | 0.996996 |

| Location | 19,607,428 – 19,607,531 |
|---|---|
| Length | 103 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 76.29 |
| Mean single sequence MFE | -23.76 |
| Consensus MFE | -8.00 |
| Energy contribution | -8.52 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.34 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.915607 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19607428 103 + 23771897 UCAUAAUUCAUAAUUCCUUCUAGUAUUGCUUUGAAAUGCUUUUAGGAAGUCCUUCAAA-UUUGAUUUCGUACAUAU--GUAUAUGUACAUAUGCGUGUGU-----AUGCGC-------- (((..........(((((...((((((.......))))))...)))))..........-..)))...(((((((((--((((((....))))))))))))-----)))...-------- ( -27.15) >DroSec_CAF1 22444 103 + 1 UCAUAAUUCAUAAUUCCUUCUAGUAUUGCUUUGAAAUGCUUUCAGGAAGUCCUUCAAA-UUUGAUUUAGUACAUAU--GUAUAUGUACAUAUACGAGUGU-----AUGUGC-------- (((..........(((((...((((((.......))))))...)))))..........-..)))....((((((.(--((((((....))))))).))))-----))....-------- ( -20.95) >DroSim_CAF1 22289 103 + 1 UCAUAAUUCAUAAUUCCUUCUAGUAUUGCUUUGAAAUGCUUUCAGGAAGUCCUUCAAA-UUUGAUUGAGUACAUAU--GUAUAUGUACAUAUACGUGUGU-----AUGUGC-------- .............(((((...((((((.......))))))...))))).((..(((..-..)))..))((((((((--((((((....))))))))))))-----))....-------- ( -26.10) >DroEre_CAF1 22676 116 + 1 ACAUAA-UCACAAUUGCAUCUAGAAUGGCUUUGAAAUGGUUUUAGGAAGUCCUUCAAAAUGAUAUUCCAUACACAU--GUAUAUGUACUUAUGCUCGUAUAGUGCAUGUAUAUUAAAUA ......-...............(((.((((((.(((....)))..)))))).)))...................((--(((((((((((.(((....))))))))))))))))...... ( -21.00) >DroYak_CAF1 22172 111 + 1 GCAUAA-UCACAAUUGCAUCUAGAAUUGCUUUGAAAUAGUUUAAGGAAGUCCUUCAAAAUCAGAUUCCAUACACAAGUGUAUAUUUAUGUGUGCAC-------GCAUGUGUAUUAUACA (.((((-(((((..(((.....(((((..(((((.........(((....))))))))....))))).........((((((((....))))))))-------))))))).))))).). ( -23.60) >consensus UCAUAAUUCAUAAUUCCUUCUAGUAUUGCUUUGAAAUGCUUUCAGGAAGUCCUUCAAA_UUUGAUUCCGUACAUAU__GUAUAUGUACAUAUGCGCGUGU_____AUGUGC________ ........((((..........((((((.((((((..(((((...)))))..))))))........))))))......((((((....))))))............))))......... ( -8.00 = -8.52 + 0.52)

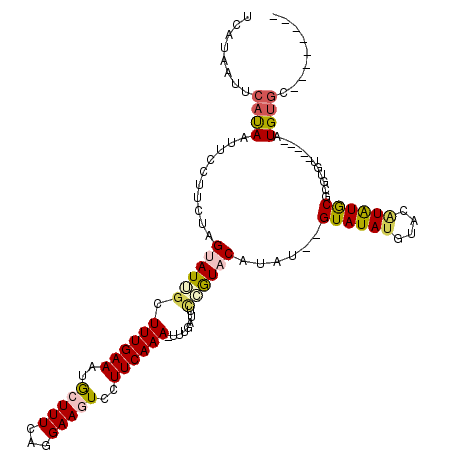

| Location | 19,607,467 – 19,607,561 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 72.64 |
| Mean single sequence MFE | -27.90 |
| Consensus MFE | -11.73 |
| Energy contribution | -11.79 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.80 |
| Structure conservation index | 0.42 |
| SVM decision value | 1.38 |
| SVM RNA-class probability | 0.948134 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19607467 94 + 23771897 UUUUAGGAAGUCCUUCAAA-UUUGAUUUCGUACAUAU--GUAUAUGUACAUAUGCGUGUGU--AUGCGC------------------UCUAGGAAUAGUGCAAGUGUAUCCUAAAGC ((((((((.(..(((....-........(((((((((--((((((....))))))))))))--))).((------------------.(((....))).)))))..).)))))))). ( -29.90) >DroSec_CAF1 22483 95 + 1 UUUCAGGAAGUCCUUCAAA-UUUGAUUUAGUACAUAU--GUAUAUGUACAUAUACGAGUGU--AUGUGC-----------------GUCUAGGAGUAGUGCAAGUGAAUCCUAGAAC .(((((((..(((((((..-..)))....((((((.(--.((.(((((((((((....)))--))))))-----------------)).)).).)).)))).)).)).)))).))). ( -22.20) >DroSim_CAF1 22328 95 + 1 UUUCAGGAAGUCCUUCAAA-UUUGAUUGAGUACAUAU--GUAUAUGUACAUAUACGUGUGU--AUGUGC-----------------CUCUAGGACUAGUGCAAGUGAUUCCUAGAAC .(((((((((((((((((.-.....))))((((((((--((((((....))))))))))))--))....-----------------....))))))............)))).))). ( -26.90) >DroYak_CAF1 22210 113 + 1 UUUAAGGAAGUCCUUCAAAAUCAGAUUCCAUACACAAGUGUAUAUUUAUGUGUGCAC----GCAUGUGUAUUAUACACAUGCAUUGCUCUAGGAUUAGUGCAAUUGCAUCCGGCAGC .....(((((((((.(((...................((((((((....))))))))----(((((((((...))))))))).)))....))))))..(((....))))))...... ( -32.60) >consensus UUUCAGGAAGUCCUUCAAA_UUUGAUUUAGUACAUAU__GUAUAUGUACAUAUACGUGUGU__AUGUGC_________________CUCUAGGAAUAGUGCAAGUGAAUCCUAGAAC ....(((....)))...............((((((....((((((....))))))........))))))..................(((((((..............))))))).. (-11.73 = -11.79 + 0.06)

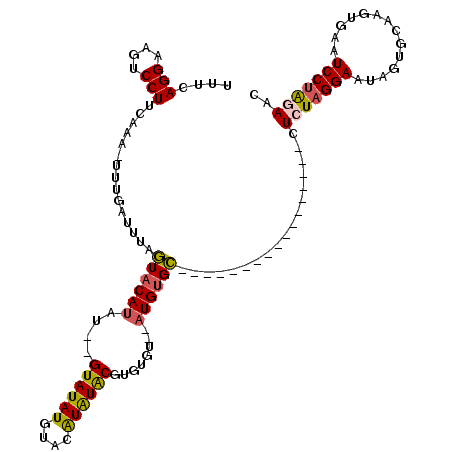
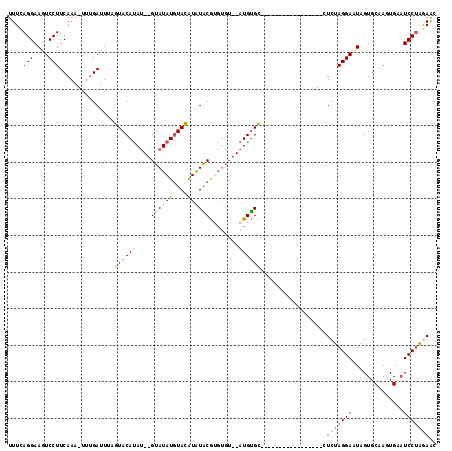
| Location | 19,607,467 – 19,607,561 |
|---|---|
| Length | 94 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 72.64 |
| Mean single sequence MFE | -23.82 |
| Consensus MFE | -11.83 |
| Energy contribution | -12.14 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.89 |
| Structure conservation index | 0.50 |
| SVM decision value | 2.78 |
| SVM RNA-class probability | 0.996996 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19607467 94 - 23771897 GCUUUAGGAUACACUUGCACUAUUCCUAGA------------------GCGCAU--ACACACGCAUAUGUACAUAUAC--AUAUGUACGAAAUCAAA-UUUGAAGGACUUCCUAAAA (((((((((..............)))))))------------------))....--......(((((((((....)))--)))))).((((......-)))).(((....))).... ( -23.64) >DroSec_CAF1 22483 95 - 1 GUUCUAGGAUUCACUUGCACUACUCCUAGAC-----------------GCACAU--ACACUCGUAUAUGUACAUAUAC--AUAUGUACUAAAUCAAA-UUUGAAGGACUUCCUGAAA (((((((((..............))))))).-----------------))..((--((....))))..(((((((...--.))))))).........-.....(((....))).... ( -17.54) >DroSim_CAF1 22328 95 - 1 GUUCUAGGAAUCACUUGCACUAGUCCUAGAG-----------------GCACAU--ACACACGUAUAUGUACAUAUAC--AUAUGUACUCAAUCAAA-UUUGAAGGACUUCCUGAAA .(((.((((............((((((.(((-----------------..((((--(.....((((((....))))))--.))))).)))..(((..-..)))))))))))))))). ( -21.20) >DroYak_CAF1 22210 113 - 1 GCUGCCGGAUGCAAUUGCACUAAUCCUAGAGCAAUGCAUGUGUAUAAUACACAUGC----GUGCACACAUAAAUAUACACUUGUGUAUGGAAUCUGAUUUUGAAGGACUUCCUUAAA .....(((((((((((((.(((....))).)))))(((((((((...)))))))))----.))).........((((((....))))))..)))))......((((....))))... ( -32.90) >consensus GCUCUAGGAUUCACUUGCACUAAUCCUAGAG_________________GCACAU__ACACACGCAUAUGUACAUAUAC__AUAUGUACGAAAUCAAA_UUUGAAGGACUUCCUGAAA ..(((((((..............)))))))......................................(((((((......)))))))...............(((....))).... (-11.83 = -12.14 + 0.31)


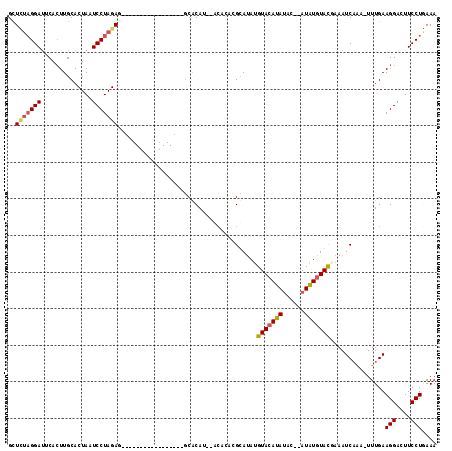
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:43:52 2006