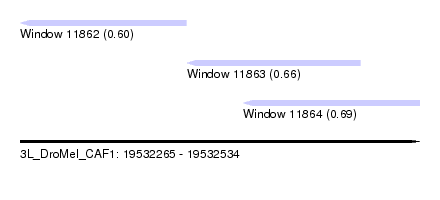
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,532,265 – 19,532,534 |
| Length | 269 |
| Max. P | 0.693766 |

| Location | 19,532,265 – 19,532,377 |
|---|---|
| Length | 112 |
| Sequences | 3 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 96.43 |
| Mean single sequence MFE | -22.41 |
| Consensus MFE | -21.12 |
| Energy contribution | -21.12 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.600371 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19532265 112 - 23771897 AAAUUUGGUUACUUAAACCAACUCUUGUAGCGAUAUUCUAAAAGAAAACCAAAACUAUCAGAUAAACGAAAGUCGGGGUCGAAACCCAAAUCCGAUGGAGGAAGACGACAUU ....((((((.....)))))).(((.((((.....(((.....)))........)))).))).........((((((((....))))...(((......)))...))))... ( -21.02) >DroSec_CAF1 2506 112 - 1 AAAUUUGGUUACUAAAACCAACUCUUGUAGCGAUAUUCUAAAAGAAAACCAAAACUAUCAGAUUAACGAAAGUCGGGGUCGAAACCCAAAUCCGAUGGAGGAAGACGACAUU ....((((((.....)))))).((((.(((.......))).)))).........(((((.((((.((....))..((((....)))).)))).))))).............. ( -21.50) >DroYak_CAF1 15977 111 - 1 AAAUUUGGUUACUAAAACCAACUCUUGUAGCGAUAUUCUAAAAGAAAACCA-AUCUAUCAGAUAAACGAAAGUCGGGGUAGAAACCCAUAUCCGAUGGAGGAAGACGACAUU ....((((((.....)))))).((((.(((.......))).))))...((.-.((((((.((((.((....))..((((....)))).)))).))))))))........... ( -24.70) >consensus AAAUUUGGUUACUAAAACCAACUCUUGUAGCGAUAUUCUAAAAGAAAACCAAAACUAUCAGAUAAACGAAAGUCGGGGUCGAAACCCAAAUCCGAUGGAGGAAGACGACAUU ....((((((.....)))))).(((.((((.....(((.....)))........)))).))).........((((((((....))))...(((......)))...))))... (-21.12 = -21.12 + -0.00)


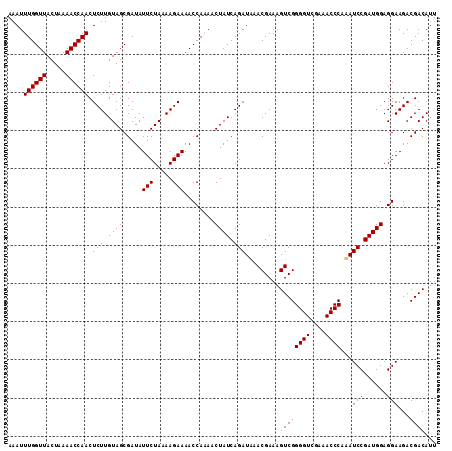
| Location | 19,532,377 – 19,532,494 |
|---|---|
| Length | 117 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.99 |
| Mean single sequence MFE | -21.64 |
| Consensus MFE | -16.86 |
| Energy contribution | -17.53 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.661423 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
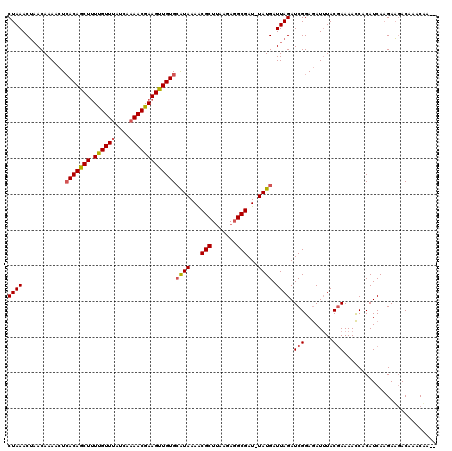
>3L_DroMel_CAF1 19532377 117 - 23771897 CUAAACUAACAAAACUCACAGCUUUUGUUUAUCAAAACGAAGUUGUGCAUAAAACGCUUAAGAUGCGAU-UAUGAUUAGAUCGGAGAUUUACGAAAACCACAUCAAGAAGACAAACUA-- .....((((.......(((((((((.((((....)))))))))))))(((((..(((.......))).)-)))).)))).(((........)))........................-- ( -20.00) >DroSec_CAF1 2618 120 - 1 CUAAACAAACAAAACUAACAGCUUUUGUUUAUCAAAACGAAGUUGUGCAUAAAGCGCUUAAGAGGCGAUUUAUGAUUAGAUCGGAGAUUUACGAAAACCACAUCACGAUAACAAACAAAG ((((.............((((((((.((((....)))))))))))).((((((.((((.....)))).)))))).))))((((..(((.............))).))))........... ( -22.02) >DroYak_CAF1 16088 117 - 1 CUAAACUUACAAAAAUCACAGCUGUCGUUAAUCAAAACGAAGCUGUGAGUAAAACGCUAAAGAGGCGAU-UAUGAUUAGAUCGGAAAUAUACAAAAAUCACAUCAAGAAGACAAGCAA-- ((((...........((((((((.(((((......)))))))))))))......((((.....))))..-.....)))).......................................-- ( -22.90) >consensus CUAAACUAACAAAACUCACAGCUUUUGUUUAUCAAAACGAAGUUGUGCAUAAAACGCUUAAGAGGCGAU_UAUGAUUAGAUCGGAGAUUUACGAAAACCACAUCAAGAAGACAAACAA__ ((((............(((((((.((((((....)))))))))))))((((...(((.......)))...)))).)))).(((........))).......................... (-16.86 = -17.53 + 0.67)


| Location | 19,532,415 – 19,532,534 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.69 |
| Mean single sequence MFE | -23.41 |
| Consensus MFE | -17.99 |
| Energy contribution | -18.33 |
| Covariance contribution | 0.34 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.41 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.693766 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
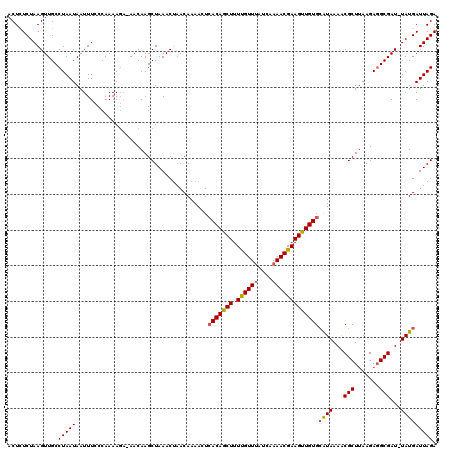
>3L_DroMel_CAF1 19532415 119 - 23771897 ACUCUCUAAGUUGCCUAAUAAUUUCCCAAAAGAUAACAAGCUAAACUAACAAAACUCACAGCUUUUGUUUAUCAAAACGAAGUUGUGCAUAAAACGCUUAAGAUGCGAU-UAUGAUUAGA ..............(((((.....................................(((((((((.((((....)))))))))))))(((((..(((.......))).)-))))))))). ( -19.00) >DroSec_CAF1 2658 120 - 1 ACUCUCUAAGUUGCCUAAUAAUUUCCCAAAACAAAACAAGCUAAACAAACAAAACUAACAGCUUUUGUUUAUCAAAACGAAGUUGUGCAUAAAGCGCUUAAGAGGCGAUUUAUGAUUAGA ....((((((((((((...((((((........(((((((((.................))))..)))))........))))))((((.....)))).....)))))))))).))..... ( -23.02) >DroYak_CAF1 16126 117 - 1 ACUUUCUAAGUUGCCUAAUAAUUUCCCAAAAG--AACAAGCUAAACUUACAAAAAUCACAGCUGUCGUUAAUCAAAACGAAGCUGUGAGUAAAACGCUAAAGAGGCGAU-UAUGAUUAGA .((.((..((((((((......(((......)--))..(((.....((((.....((((((((.(((((......)))))))))))))))))...)))....)))))))-)..))..)). ( -28.20) >consensus ACUCUCUAAGUUGCCUAAUAAUUUCCCAAAAGA_AACAAGCUAAACUAACAAAACUCACAGCUUUUGUUUAUCAAAACGAAGUUGUGCAUAAAACGCUUAAGAGGCGAU_UAUGAUUAGA ..............(((((.....................................(((((((.((((((....)))))))))))))((((...(((.......)))...))))))))). (-17.99 = -18.33 + 0.34)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:43:27 2006