
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,218,499 – 19,218,750 |
| Length | 251 |
| Max. P | 0.990801 |

| Location | 19,218,499 – 19,218,598 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 99.33 |
| Mean single sequence MFE | -24.10 |
| Consensus MFE | -24.10 |
| Energy contribution | -24.10 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.89 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.07 |
| SVM RNA-class probability | 0.987187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19218499 99 + 23771897 GGAAAGGCCAAACUAGAAAAUGGGGCCAGGGAAAAGCAAAAGCAGCUUUACCAGGCAAAGGCUAAAAAGUAAACGCCUUUGGUUAGUGAAAAGUGAAAG .....((((...(........).))))..((.(((((.......))))).))((.(((((((............))))))).))............... ( -24.10) >DroSec_CAF1 47234 99 + 1 GGAAAGGCCAAACUGGAAAAUGGGGCCAGGGAAAAGCAAAAGCAGCUUUACCAGGCAAAGGCUAAAAAGUAAACGCCUUUGGUUAGUGAAAAGUGAAAG .....((((...(........).))))..((.(((((.......))))).))((.(((((((............))))))).))............... ( -24.10) >DroSim_CAF1 50987 99 + 1 GGAAAGGCCAAACUGGAAAAUGGGGCCAGGGAAAAGCAAAAGCAGCUUUACCAGGCAAAGGCUAAAAAGUAAACGCCUUUGGUUAGUGAAAAGUGAAAG .....((((...(........).))))..((.(((((.......))))).))((.(((((((............))))))).))............... ( -24.10) >consensus GGAAAGGCCAAACUGGAAAAUGGGGCCAGGGAAAAGCAAAAGCAGCUUUACCAGGCAAAGGCUAAAAAGUAAACGCCUUUGGUUAGUGAAAAGUGAAAG .....((((...(........).))))..((.(((((.......))))).))((.(((((((............))))))).))............... (-24.10 = -24.10 + -0.00)


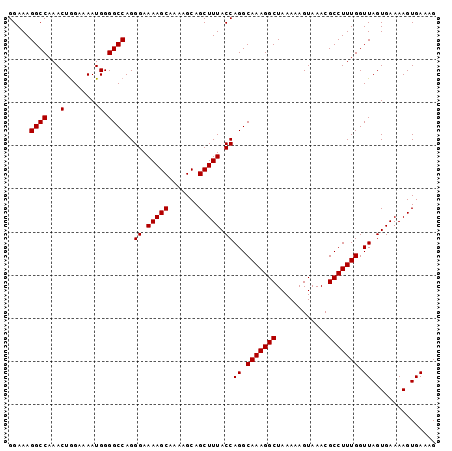
| Location | 19,218,499 – 19,218,598 |
|---|---|
| Length | 99 |
| Sequences | 3 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 99.33 |
| Mean single sequence MFE | -21.40 |
| Consensus MFE | -21.33 |
| Energy contribution | -21.33 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.28 |
| Structure conservation index | 1.00 |
| SVM decision value | 2.23 |
| SVM RNA-class probability | 0.990801 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19218499 99 - 23771897 CUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGCUGCUUUUGCUUUUCCCUGGCCCCAUUUUCUAGUUUGGCCUUUCC ............((((..(((((((............)))))))..))))(((((.......))))).....((((..(((....)))..))))..... ( -21.80) >DroSec_CAF1 47234 99 - 1 CUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGCUGCUUUUGCUUUUCCCUGGCCCCAUUUUCCAGUUUGGCCUUUCC ............((((..(((((((............)))))))..))))(((((.......))))).....((((..((......))..))))..... ( -21.20) >DroSim_CAF1 50987 99 - 1 CUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGCUGCUUUUGCUUUUCCCUGGCCCCAUUUUCCAGUUUGGCCUUUCC ............((((..(((((((............)))))))..))))(((((.......))))).....((((..((......))..))))..... ( -21.20) >consensus CUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGCUGCUUUUGCUUUUCCCUGGCCCCAUUUUCCAGUUUGGCCUUUCC ............((((..(((((((............)))))))..))))(((((.......))))).....((((..(((....)))..))))..... (-21.33 = -21.33 + -0.00)



| Location | 19,218,539 – 19,218,638 |
|---|---|
| Length | 99 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 88.53 |
| Mean single sequence MFE | -25.83 |
| Consensus MFE | -22.52 |
| Energy contribution | -22.15 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.533954 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19218539 99 - 23771897 UGCUCCCGAGUCGGCGGCUGUCGUAUUUGUUUACGCUGCUCUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGC--------------------UGCU .(((...(((..((((((....(((......)))))))))..))).......((((..(((((((............)))))))..))))..)))--------------------.... ( -25.60) >DroSec_CAF1 47274 99 - 1 UGCUCCCGAGUCGGUGGCUGUCGUAUUUGUUUACGCUGCUCUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGC--------------------UGCU .(((...(((.(((((.....((....))....))))))))...........((((..(((((((............)))))))..))))..)))--------------------.... ( -23.10) >DroSim_CAF1 51027 99 - 1 UGCUCCCGAGUCGGCGGCUGUCGUAUUUGUUUACGCUGCUCUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGC--------------------UGCU .(((...(((..((((((....(((......)))))))))..))).......((((..(((((((............)))))))..))))..)))--------------------.... ( -25.60) >DroYak_CAF1 76249 119 - 1 UGGUCCCGAGUCGGCGACUCUUGUAUUUGUUUACGCUGCUCUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGCUUGGCUUUUGCUUCGGCUUCUGCU .......(((.(((((...(........)....)))))))).....((((..((((..(((((((............)))))))..))))))))...(((....((....))....))) ( -29.00) >consensus UGCUCCCGAGUCGGCGGCUGUCGUAUUUGUUUACGCUGCUCUUUCACUUUUCACUAACCAAAGGCGUUUACUUUUUAGCCUUUGCCUGGUAAAGC____________________UGCU .......(((.(((((.....((....))....)))))))).....((((..((((..(((((((............)))))))..))))))))......................... (-22.52 = -22.15 + -0.37)



| Location | 19,218,638 – 19,218,750 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 85.55 |
| Mean single sequence MFE | -38.02 |
| Consensus MFE | -26.88 |
| Energy contribution | -26.92 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.582511 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19218638 112 + 23771897 ACUUACAUCUGCUGAUAAAGAAAACAGCAGGCAUCCCAUGUGAGCCAUGUCCAUAGGUUAUAAAUCGUGGUCGGGCUG-----CCUGGUUGGGUGGUUCCGACCUGGCUAGGUAGUU--- .(((((((((((((..........)))))).......)))))))...((.((((.(((.....))))))).))(((((-----((((((..((((....).)))..)))))))))))--- ( -40.81) >DroSec_CAF1 47373 110 + 1 ACUUACAUCUGCUGAUAAAGAAAACAGCAGGCAUCCCAUGUGAGCCACGUCCAUAGGUUAUAAAUCGUGGUUGGGCUG-----CCUGGUUGGGUGGUUCCGACCUGGCUAGGUUG----- ........((((((..........))))))(((.((((.....((((((................)))))))))).))-----)(((((..((((....).)))..)))))....----- ( -37.59) >DroSim_CAF1 51126 110 + 1 ACUUACAUCUGCUGAUAAAGAAAACAGCAGGCAUCCCAUGUGAGCCAUGUCCAUAGGUUAUAAAUCGUGGUUGGGCUG-----CCUGGUUGGGUGGUUCCGACCUGGCUAGGUUG----- .(((((((((((((..........)))))).......)))))))(((...((((.(((.....))))))).)))...(-----((((((..((((....).)))..)))))))..----- ( -36.81) >DroEre_CAF1 49643 119 + 1 ACUUACAUCUGCUGAUAAAGAAAACAGCAGGCAUCCUAUGUGAGCCAUGUUCAUAGUUUAUAAAUCGUGGUUGGGUUGGUUAGGCCGGUGGGGCGGUUCCGACCUGGCUGAGUGGGUGG- .......(((((((..........)))))))(((((..(((((((...))))))).............(((..((((((....(((.....)))....))))))..)))....))))).- ( -40.60) >DroYak_CAF1 76368 106 + 1 ACUUACAUCUCCUGAUAAAGAAAACUGCAGGCAUCCUAUGUGAGCCAUG----UAGUUUAUAAAUCAUGGUUG----------GCUGGUUGGGCGGUUUCGACCUGGCUGGGUGGGUGCU ((((((....((((((.....(((((((((((.((......))))).))----))))))....)))).))...----------.(..((..((((....)).))..))..)))))))... ( -34.30) >consensus ACUUACAUCUGCUGAUAAAGAAAACAGCAGGCAUCCCAUGUGAGCCAUGUCCAUAGGUUAUAAAUCGUGGUUGGGCUG_____CCUGGUUGGGUGGUUCCGACCUGGCUAGGUGG_U___ .(((((((((((((..........)))))).......)))))))(((((................))))).............((((((..((((....)).))..))))))........ (-26.88 = -26.92 + 0.04)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:39:49 2006