
| Sequence ID | 3L_DroMel_CAF1 |
|---|---|
| Location | 19,190,678 – 19,190,793 |
| Length | 115 |
| Max. P | 0.999942 |

| Location | 19,190,678 – 19,190,793 |
|---|---|
| Length | 115 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 69.32 |
| Mean single sequence MFE | -20.79 |
| Consensus MFE | -14.88 |
| Energy contribution | -14.00 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.72 |
| SVM decision value | 4.71 |
| SVM RNA-class probability | 0.999942 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
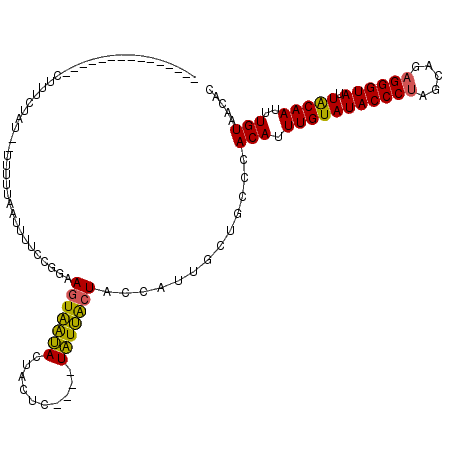
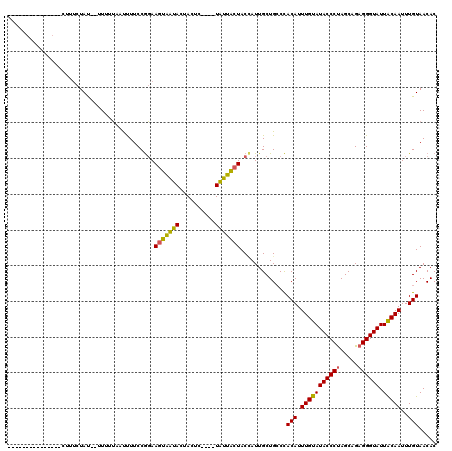
>3L_DroMel_CAF1 19190678 115 + 23771897 AUUUUCCGGAAGUAA-----UACUACUCUACCCCUUCCAUAAUUCGAAUAACUUUUGGUUUGGCUACUAUUGCUGCCUACAUUUGUAUACCCCAGCAGAGGGUAUUACAAUUUGUAACAC .......((.(((((-----((.((((..(((........................)))..)).)).))))))).))((((.(((((((((((....).))))).)))))..)))).... ( -22.56) >DroSec_CAF1 19077 99 + 1 ---------------CUUUCUAU--UUUUUUAUUUUCCGGAAGUAAUACUACUC----UAUUACUACCAUUGCUGGCCACAUUUGUAUACCCUAGCAGAGGGUAUUACAAUUUGUAACAC ---------------........--.............((.(((((((......----))))))).))..........(((.(((((((((((.....)))))).)))))..)))..... ( -21.00) >DroSim_CAF1 23369 99 + 1 ---------------CUUUCUAU--UUUUUAAUUUUCCGGAAGUAAUACUACUC----UAUUACUACGAUUGCUGCCCACAUUUGUAUACCCUAGCAGAGGGUAUUGCAAUUUGUAACAC ---------------........--............((..(((((((......----))))))).))..........(((.(((((((((((.....)))))).)))))..)))..... ( -18.80) >consensus _______________CUUUCUAU__UUUUUAAUUUUCCGGAAGUAAUACUACUC____UAUUACUACCAUUGCUGCCCACAUUUGUAUACCCUAGCAGAGGGUAUUACAAUUUGUAACAC .........................................(((((((..........))))))).............(((.(((((((((((.....)))))).)))))..)))..... (-14.88 = -14.00 + -0.88)

| Location | 19,190,678 – 19,190,793 |
|---|---|
| Length | 115 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.32 |
| Mean single sequence MFE | -23.07 |
| Consensus MFE | -13.08 |
| Energy contribution | -13.53 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.82 |
| SVM RNA-class probability | 0.858361 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>3L_DroMel_CAF1 19190678 115 - 23771897 GUGUUACAAAUUGUAAUACCCUCUGCUGGGGUAUACAAAUGUAGGCAGCAAUAGUAGCCAAACCAAAAGUUAUUCGAAUUAUGGAAGGGGUAGAGUAGUA-----UUACUUCCGGAAAAU ....((((..(((((.((((((.....))))))))))).))))(((.((....)).)))...((...((((.....))))..(((((..(((......))-----)..)))))))..... ( -28.00) >DroSec_CAF1 19077 99 - 1 GUGUUACAAAUUGUAAUACCCUCUGCUAGGGUAUACAAAUGUGGCCAGCAAUGGUAGUAAUA----GAGUAGUAUUACUUCCGGAAAAUAAAAAA--AUAGAAAG--------------- (.((((((..(((((.((((((.....))))))))))).))))))).....(((.(((((((----......))))))).)))............--........--------------- ( -23.90) >DroSim_CAF1 23369 99 - 1 GUGUUACAAAUUGCAAUACCCUCUGCUAGGGUAUACAAAUGUGGGCAGCAAUCGUAGUAAUA----GAGUAGUAUUACUUCCGGAAAAUUAAAAA--AUAGAAAG--------------- ..........((((.(((((((.....))))))).....((....))))))(((.(((((((----......)))))))..)))...........--........--------------- ( -17.30) >consensus GUGUUACAAAUUGUAAUACCCUCUGCUAGGGUAUACAAAUGUGGGCAGCAAUAGUAGUAAUA____GAGUAGUAUUACUUCCGGAAAAUAAAAAA__AUAGAAAG_______________ .(((((((..(((((.((((((.....))))))))))).))))).))....(((.(((((((..........))))))).)))..................................... (-13.08 = -13.53 + 0.45)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:39:11 2006